Mastering Error-Prone PCR: Strategies to Overcome Mutagenesis Bias for Directed Evolution Success
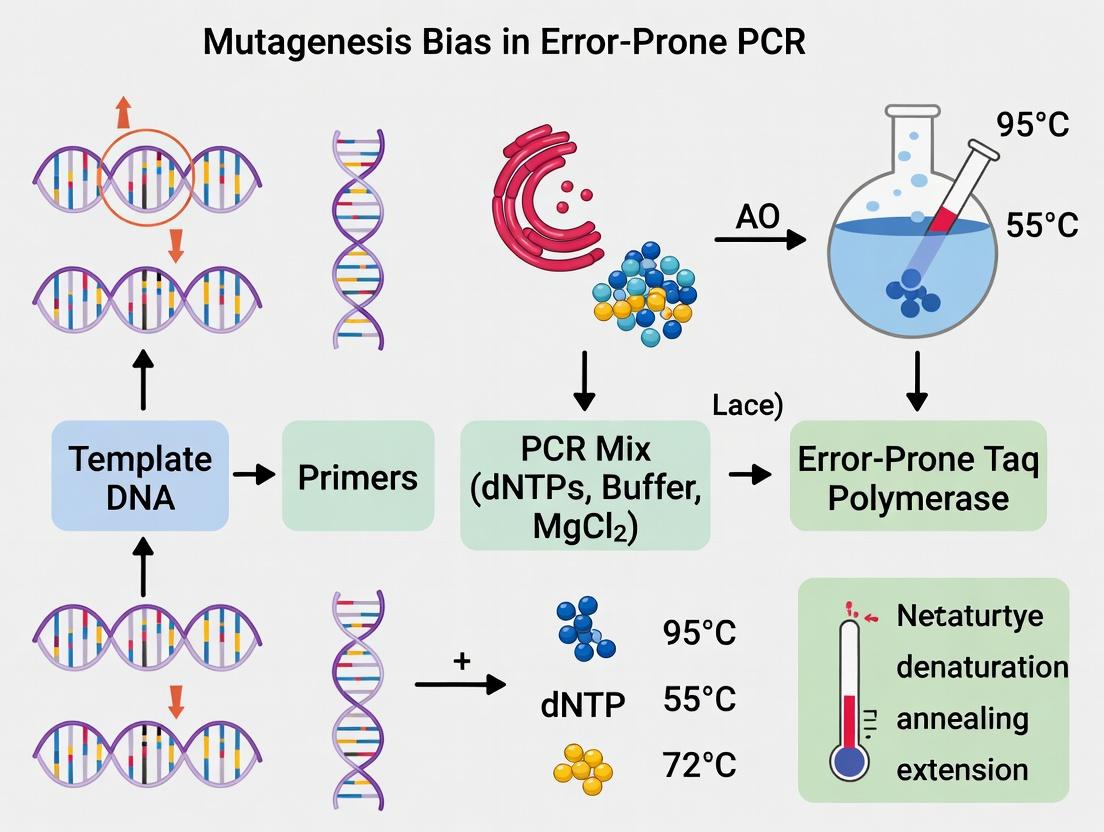
This article provides a comprehensive guide for researchers on understanding and mitigating the intrinsic biases of error-prone PCR (epPCR) in directed evolution and protein engineering.
Mastering Error-Prone PCR: Strategies to Overcome Mutagenesis Bias for Directed Evolution Success
Abstract
This article provides a comprehensive guide for researchers on understanding and mitigating the intrinsic biases of error-prone PCR (epPCR) in directed evolution and protein engineering. We explore the biochemical foundations of mutation bias, detail current methodological approaches to control mutational spectra, offer troubleshooting frameworks for common experimental pitfalls, and present comparative validation strategies. The content equips scientists with the knowledge to design robust epPCR protocols that generate high-quality, diverse mutant libraries, directly impacting the efficiency of discovering novel enzymes, antibodies, and therapeutic proteins.
The Roots of Randomness: Understanding the Inherent Biases in Error-Prone PCR
Technical Support Center: Troubleshooting Unbiased Mutational Libraries
FAQs & Troubleshooting Guides
Q1: My error-prone PCR (epPCR) library shows a strong nucleotide sequence bias, with over-representation of A/T to G/C transitions. What are the primary causes and solutions?
A: This is a classic symptom of manganese-induced bias in standard Taq polymerase epPCR protocols. The bias originates from Mn²⁺ ions facilitating misincorporation of certain dNTPs.
- Solution: Implement a balanced nucleotide analog protocol. Use a mutator strain like Mutazyme II or a polymerase blend designed for unbiased mutagenesis (e.g., incorporating Deep Vent or Dpo4 variants). Ensure the dNTP concentration is precisely balanced (see Table 1). A checklist is provided in Diagram 1.
Q2: The mutation rate from my epPCR is consistently too low (< 0.1% per base) for effective screening, even with adjusted Mn²⁺/Mg²⁺ ratios. How can I increase it without introducing bias?
A: Low mutation rates often stem from overly stringent reaction conditions that favor correct nucleotide incorporation.
- Solution:
- Validate Nucleotide Analogs: Use a calibrated mix of dPTP and 8-oxo-dGTP. These analogs are incorporated opposite multiple natural bases, promoting transversion mutations which are often underrepresented.
- Optimize Template: Ensure template DNA is clean and quantified accurately. Excess template dilutes mutation frequency.
- Cycle Optimization: Increase the number of PCR cycles (e.g., 30-35 cycles) to accumulate mutations. Refer to the detailed protocol in the "Experimental Protocols" section.
Q3: How can I quantitatively assess the bias and quality of my generated mutant library before proceeding to screening?
A: Pre-screening validation is critical. Perform Next-Generation Sequencing (NGS) on a representative sample of your library (minimum 10⁴ clones).
- Analysis: Calculate the observed frequency of each possible nucleotide substitution (A→T, A→C, A→G, etc.). Compare this to the expected frequency under a perfectly random model (6.25% for each of the 12 possible substitution types given a base). Use the normalized entropy score (see Table 2). A score closer to 1 indicates lower bias.
Data Presentation
Table 1: Comparison of Key Unbiased Mutagenesis Methods & Their Performance Metrics
| Method | Core Mechanism | Typical Mutation Rate (%/base) | Transition:Transversion Ratio (Ideal = 1:2) | Key Advantage | Primary Bias Risk |
|---|---|---|---|---|---|
| Standard Mn²⁺/Mg²⁺ epPCR | Mn²⁺ reduces fidelity of Taq pol. | 0.1 - 0.5 | ~4:1 to 10:1 | Simple, low-cost | High AT→GC bias |
| Nucleotide Analog (dPTP/8-oxo-dGTP) | Base analogs pair with multiple partners. | 0.5 - 2.0 | ~1.5:1 to 2:1 | More balanced substitution spectrum | Slight GC→AT bias if unbalanced |
| Mutator Polymerase Blends | Engineered or natural low-fidelity polymerases. | 0.5 - 3.0 | ~1:1 to 1.5:1 | High mutational load, simpler setup | Sequence context dependence |
| Plasmid-based Mutagenesis (e.g., TRIDENT) | In vivo replication with impaired mismatch repair. | ~0.005 - 0.01 per generation | ~1:1 | Ultra-deep libraries, no PCR artifacts | Requires specialized E. coli strains |
Table 2: Quantitative Bias Assessment of a Hypothetical Mutant Library via NGS
| Mutation Type | Observed Frequency (%) | Expected Random Frequency (%) | Deviation (Observed - Expected) |
|---|---|---|---|
| A → T | 3.2 | 6.25 | -3.05 |
| A → C | 3.8 | 6.25 | -2.45 |
| A → G | 12.5 | 6.25 | +6.25 |
| T → A | 3.5 | 6.25 | -2.75 |
| T → C | 14.1 | 6.25 | +7.85 |
| T → G | 2.9 | 6.25 | -3.35 |
| G → A | 10.8 | 6.25 | +4.55 |
| G → T | 4.1 | 6.25 | -2.15 |
| G → C | 3.3 | 6.25 | -2.95 |
| C → A | 4.0 | 6.25 | -2.25 |
| C → G | 3.0 | 6.25 | -3.25 |
| C → T | 34.8 | 6.25 | +28.55 |
| Normalized Entropy Score (H/H_max) | 0.65 | 1.00 (Ideal) | N/A |
Experimental Protocols
Protocol: Unbiased Mutagenesis Using a Nucleotide Analog & Polymerase Blend
Objective: Generate a mutant library with a target mutation rate of 1-2 mutations/kb and a balanced substitution spectrum.
Materials: See "The Scientist's Toolkit" below.
Procedure:
- Reaction Setup: In a thin-walled PCR tube, assemble a 50 µL reaction on ice:
- Template DNA (100 ng - 1 µg, depending on length)
- 1X Mutagenesis Buffer (supplied with enzyme blend)
- dNTP Mix: 0.2 mM each of dATP, dTTP, 0.1 mM dCTP, 0.1 mM dGTP.
- Nucleotide Analog Mix: 0.05 mM dPTP, 0.05 mM 8-oxo-dGTP.
- MgSO₄ to a final concentration of 2 mM.
- Forward and Reverse Primers (0.5 µM each).
- Unbiased Mutagenesis Polymerase Blend (e.g., 1.0 unit).
- Thermocycling: Use the following program:
- Initial Denaturation: 95°C for 2 min.
- 30 Cycles:
- Denature: 95°C for 30 sec.
- Anneal: [Primer Tm -5°C] for 30 sec.
- Extend: 72°C for 1 min/kb of product.
- Final Extension: 72°C for 5 min.
- Hold at 4°C.
- Purification: Purify the PCR product using a spin column-based PCR purification kit to remove nucleotides, primers, and polymerase.
- Clone & Validate: Clone the product into your expression vector using your preferred method (e.g., Gibson assembly, restriction digest/ligation). Sequence 20-50 random clones via Sanger sequencing to estimate the average mutation rate and check for bias before large-scale transformation and NGS analysis.
Mandatory Visualization
Diagram 1: Troubleshooting Workflow for Biased Mutagenesis Libraries
Diagram 2: Logic of Unbiased vs. Biased Mutational Search Space
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Unbiased Mutagenesis | Example/Notes |
|---|---|---|
| Unbiased Polymerase Blends | Engineered mixtures that promote random misincorporation with minimal sequence bias. | Mutazyme II (Agilent), GeneMorph II (Agilent), commercial "low-fidelity" blends. |
| Nucleotide Analogs (dPTP, 8-oxo-dGTP) | Partially degenerate bases that pair with multiple natural bases, increasing transversion frequency. | Critical for balancing the transition:transversion ratio. Handle as light-sensitive. |
| Optimized Mutagenesis Buffers | Provide optimal ionic conditions (Mg²⁺, K⁺, etc.) for the mutator polymerase, often with Mn²⁺ excluded. | Use the buffer supplied with the enzyme blend; do not substitute. |
| High-Purity dNTP Solutions | Pre-mixed, balanced dNTP sets at low concentrations to promote misincorporation. | Avoid home-made mixes from high-concentration stocks to prevent pipetting errors. |
| PCR Purification Kits | Remove nucleotides, primers, and enzyme post-PCR to prepare clean DNA for cloning. | Essential step before library construction to prevent carryover of analogs/polymerase. |
| NGS Library Prep Kit | For deep sequencing analysis of library diversity and bias quantification. | Amplicon-based kits (e.g., Illumina) are suitable for ~500bp target regions. |
Troubleshooting Guides & FAQs
Q1: During error-prone PCR, I am observing a much lower mutation frequency than expected. What are the primary causes? A1: Low mutation frequency typically stems from insufficient error-inducing conditions. First, verify the concentration of MnCl₂ in your reaction; manganese is crucial for reducing polymerase fidelity. Ensure it is between 0.1-0.5 mM. Second, check the dNTP imbalance. A standard protocol uses an unequal dNTP pool (e.g., increasing [dATP] and [dTTP] while decreasing [dCTP] and [dGTP]) to promote misincorporation. Third, confirm you are using a polymerase with no 3'→5' exonuclease proofreading activity (e.g., Taq polymerase). Using a high-fidelity polymerase will correct errors.
Q2: My mutation spectrum is skewed heavily toward specific transversions. How can I achieve a more random/balanced spectrum? A2: A skewed spectrum often indicates an overly aggressive dNTP imbalance. To promote randomness, adjust your dNTP ratios closer to equimolar while maintaining an overall elevated dNTP concentration (e.g., 0.2 mM each dNTP). Additionally, supplementing with nucleoside analogs like 8-oxo-dGTP or dPTP can diversify mutation types. Ensure Mg²⁺ and Mn²⁺ concentrations are optimized, as their ratio directly influences polymerase error bias.
Q3: The product yield from my error-prone PCR is very poor. How can I improve amplification efficiency? A3: Poor yield is common due to suboptimal cycling conditions for error-prone mixtures. Increase the number of cycles (e.g., 30-40 cycles). Extend the extension time to accommodate potentially slower polymerization. Titrate the MnCl₂ concentration; too much Mn²⁺ can inhibit amplification. Perform a gradient PCR to optimize the annealing temperature, as the altered dNTP pool can affect primer binding. Adding DMSO (2-5%) or betaine can help stabilize the reaction.
Q4: How do I verify and quantify the mutation rate and spectrum from my library? A4: This requires sequencing. The gold standard is deep sequencing of the amplified target region from the pooled library. For a cost-effective initial assessment, clone individual products (e.g., 50-100 colonies) and perform Sanger sequencing. Calculate the mutation rate as (total mutations) / (total base pairs sequenced). Analyze the spectrum by categorizing mutations (A→G, A→T, etc.).
Q5: What controls are essential for a reliable error-prone PCR experiment? A5: Always run these controls in parallel: 1) Standard PCR Control: A reaction with standard, balanced dNTPs and no Mn²⁺ to confirm template integrity and baseline amplification. 2) Error-Prone No-Template Control: To check for contamination. 3) Replicate Reactions: Error-prone PCR is stochastic; perform at least 3 replicate library constructions to ensure reproducibility.
Key Quantitative Data for Error-Prone PCR Optimization
Table 1: Effects of Mn²⁺ Concentration on Mutation Frequency (Using Taq Polymerase)
| [MnCl₂] (mM) | Relative Mutation Frequency (per kb) | Primary Effect on Spectrum |
|---|---|---|
| 0.0 | 0.5 - 2 | Baseline PCR errors |
| 0.1 | 5 - 10 | Slight increase in transversions |
| 0.3 | 10 - 30 | Broad increase, more random |
| 0.5 | 30 - 50 | Very high, potential bias & yield loss |
| 0.7 | Often inhibits amplification | N/A |
Table 2: Common dNTP Imbalance Protocols for Skewing Mutation Spectra
| Desired Bias | Example dNTP Ratios (A:T:C:G in mM) | Typical Mutation Bias Induced |
|---|---|---|
| AT-rich outcomes | 1.0 : 1.0 : 0.2 : 0.2 | Increases A/T content; G/C → A/T transitions |
| Increased Transversions | 0.2 : 1.0 : 1.0 : 0.2 | Favors purine → pyrimidine changes |
| "Randomized" Library | 0.5 : 0.5 : 0.5 : 0.5 (high total [dNTP]=2mM) | More even distribution of mutation types |
Experimental Protocol: Standard Error-Prone PCR for Library Generation
Objective: To create a diverse mutant library of a 1kb gene fragment with a target mutation frequency of 5-15 mutations/kb.
Materials: See "Research Reagent Solutions" below.
Method:
- Prepare 50µL Reaction Mix:
- 1X Standard Taq Reaction Buffer
- Template DNA: 10-100 ng (plasmid or product)
- Forward & Reverse Primers: 0.5 µM each
- dNTP Mix: Prepare an unbalanced stock. For a more random spectrum, use: 0.2 mM dATP, 0.2 mM dGTP, 1.0 mM dCTP, 1.0 mM dTTP.
- MgCl₂: 2.0 mM (from buffer, may need supplemental)
- MnCl₂: 0.25 mM (add from a fresh 10 mM stock).
- Taq DNA Polymerase: 2.5 units.
- Nuclease-free water to 50 µL.
- Thermal Cycling:
- Initial Denaturation: 95°C for 3 min.
- 30 Cycles of:
- Denaturation: 95°C for 45 sec.
- Annealing: 55-60°C (optimize for primers) for 60 sec.
- Extension: 72°C for 90 sec/kb.
- Final Extension: 72°C for 5 min.
- Purification: Run the entire product on an agarose gel. Excise and purify the correctly sized band using a gel extraction kit.
- Analysis: Clone into your desired vector and sequence a representative number of clones (e.g., 10-12) to empirically determine the mutation rate and spectrum before proceeding with library construction.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Error-Prone PCR Studies
| Reagent/Material | Function & Critical Notes |
|---|---|
| Taq DNA Polymerase | Lacks 3'→5' proofreading activity, essential for retaining incorporated errors. Do not use high-fidelity polymerases. |
| Manganese Chloride (MnCl₂) | The key fidelity-disruptor. Reduces base-pairing stringency. Must be titrated (0.1-0.5 mM). Prepare fresh stock. |
| Unbalanced dNTP Set | Individual dNTP solutions allow custom imbalanced pools to bias mutation spectra (e.g., favor transitions/transversions). |
| Nucleoside Analogs (e.g., 8-oxo-dGTP, dPTP) | Used to further diversify mutation types by providing alternative bases for incorporation during PCR. |
| Gel Extraction Kit | Critical for isolating the correctly sized error-prone PCR product from primers, primer dimers, and non-specific fragments. |
| TA Cloning Kit | For easy cloning of Taq-amplified products (with A-overhangs) for initial sequencing analysis of mutation spectrum. |
| Next-Generation Sequencing (NGS) Platform | For comprehensive, quantitative analysis of the final mutant library's complexity, mutation rate, and spectrum. |
Visualizations
Error-Prone PCR Workflow for Mutagenesis Analysis
Factors Influencing PCR Mutation Spectrum
Technical Support Center
Troubleshooting Guides & FAQs
Q1: I am using the classic method of adding Mn2+ to my PCR to introduce mutations, but my product yield is extremely low or I get no product at all. What could be wrong? A: Low yield is a common issue with Mn2+ methods. Mn2+ is a potent polymerase inhibitor and reduces fidelity/processivity. Troubleshoot as follows:
- Titrate MnCl2 concentration: Start very low (e.g., 0.01-0.05 mM) and increase incrementally (up to ~0.5 mM) alongside your standard Mg2+ concentration (e.g., 1.5 mM). Never replace Mg2+ entirely.
- Optimize dNTP ratios last: Only after establishing a yield with low Mn2+, begin imbalancing dNTPs (e.g., increasing one dNTP to 0.2-1 mM while keeping the other three at 0.05-0.1 mM).
- Check polymerase compatibility: Not all polymerases tolerate Mn2+. Use ones historically documented for mutagenesis like Taq DNA polymerase. Consider switching to a modern mutagenic polymerase designed for error-prone PCR (epPCR).
Q2: With imbalanced dNTPs, my mutation rate is too low and not diverse enough. How can I increase and diversify mutations? A: The classic dNTP imbalance method often produces biased mutational spectra (e.g., favoring A:T → G:C transitions).
- Combine methods: Use a low concentration of Mn2+ and imbalanced dNTPs. The two are synergistic for increasing error rate.
- Systematically vary the biased dNTP: Run parallel reactions where a different dNTP is in excess in each. Pool the products later for diversity.
- Shift to modern polymerases: Use engineered mutator strains (e.g., Mutazyme II) that provide more uniform mutation spectra without extreme buffer manipulation.
Q3: I switched to a commercial mutagenic polymerase kit, but the mutation frequency is still not meeting my expectations for directed evolution. How can I fine-tune it? A: Modern kits offer parameters for control.
- Use the recommended mutation rate matrix: Most kits provide guidelines (e.g., low, medium, high mutation rate) based on template amount and cycle number. Do not exceed recommended cycles.
- Adjust template quantity: Using less template (e.g., 10-100 pg) can increase the effective mutation rate per round.
- Perform sequential epPCR: Instead of one ultra-high mutation round (which risks deleterious mutations), perform 2-3 rounds of low/medium mutation rate PCR, screening/selecting between rounds.
Q4: How do I measure the actual mutation frequency and spectrum from my error-prone PCR experiment? A: You must sequence a representative sample of clones.
- Protocol: 1) Clone epPCR products into a plasmid vector. 2) Pick 10-20 random colonies for Sanger sequencing. 3) Analyze sequences against the parent gene.
- Calculation: Mutation Frequency (MF) = (Total number of mutations) / (Total number of bases sequenced). Express as % or mutations/kb.
- Spectrum Analysis: Categorize each mutation as Transition (Ti: AG, CT) or Transversion (Tv: all others) and note the base change.
Data Presentation: Comparison of Classic vs. Modern Error-Prone PCR Methods
| Parameter | Classic Method (Mn2+ + dNTP Imbalance) | Modern Method (Mutagenic Polymerase Kits) |
|---|---|---|
| Typical Mutation Rate | 0.1 - 2 mutations/kb/generation | 1 - 16+ mutations/kb/generation |
| Mutational Bias | High (Strong A:T → G:C bias) | Low-Moderate (More uniform spectrum) |
| Ease of Optimization | Difficult (Multiple interdependent variables) | Simplified (Pre-optimized buffers, clear guidelines) |
| Product Yield | Often significantly reduced | Higher, more consistent |
| Primary Control Knobs | [Mn2+], [Mg2+], individual [dNTPs], polymerase choice, cycles | Kit variant (low/High-fidelity), template amount, cycles |
| Best For | Proof-of-concept, low-rate mutagenesis, budget constraints | Directed evolution campaigns requiring controlled, higher rates |
Experimental Protocol: Standardized Error-Prone PCR using a Mutagenic Polymerase
Objective: Amplify a target gene with a controlled, low-to-medium mutation rate. Reagents: Template DNA (100 pg - 10 ng), mutagenic polymerase mix (e.g., GeneMorph II kit), forward/reverse primers, nuclease-free water. Protocol:
- Prepare 50 µL reaction: On ice, combine:
- 34.5 µL Nuclease-free water
- 10.0 µL 5X Mutagenic PCR Buffer (from kit)
- 2.0 µL Forward Primer (10 µM)
- 2.0 µL Reverse Primer (10 µM)
- 1.0 µL Template DNA (diluted to appropriate concentration)
- 0.5 µL Mutagenic Polymerase Blend
- Run Thermocycling Program:
- Initial Denaturation: 95°C for 2 min.
- Cycling (30x): 95°C for 30 sec, [TM -5°C] for 30 sec, 72°C for 1 min/kb.
- Final Extension: 72°C for 10 min.
- Hold: 4°C.
- Purify: Run product on agarose gel, excise correct band, and purify using a gel extraction kit.
- Clone & Sequence: Clone purified fragment into desired vector, transform, and sequence multiple colonies to assess mutation frequency/spectrum.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function / Role in Mutagenesis |
|---|---|
| Manganese Chloride (MnCl2) | Classic mutagenic agent. Substitutes for Mg2+ in polymerase active site, reducing fidelity. |
| Unbalanced dNTP Stock Solutions | Creating biased nucleotide pools to mislead polymerase incorporation. |
| Taq DNA Polymerase | Historically used classic polymerase with moderate tolerance for Mn2+ and mutagenic conditions. |
| Mutazyme II / GeneMorph II | Engineered mutagenic polymerase blends. Contain epPCR-optimized enzymes for higher, less biased rates. |
| High-Fidelity PCR Purification Kit | Essential for cleaning epPCR products before cloning to remove primers, salts, and polymerase. |
| Cloning Vector (e.g., blunt-end) | Vector compatible with the polishing/processing of epPCR products for library generation. |
| Competent Cells (High-Efficiency) | For transforming the mutagenized library to obtain a sufficient number of clones for screening. |
Visualizations
Diagram 1: Classic vs. Modern epPCR Workflow
Diagram 2: Mutagenesis Bias in Error-Prone PCR Methods
Troubleshooting Guides & FAQs
FAQ 1: Addressing Mutation Bias in Error-Prone PCR
Q: My error-prone PCR library shows a strong bias towards Transitions (Ts) over Transversions (Tv). How can I increase the Tv:Ts ratio to achieve a more diverse mutation spectrum? A: A skewed Ts/Tv ratio is a common issue. Implement these solutions:
- Adjust dNTP Concentrations: Lower the concentration of the dNTP corresponding to the original base (e.g., for A→G bias, lower dCTP) and increase the other three. Table 1 summarizes a recommended starting point.
- Use Modified Nucleotide Analogues: Incorporate 8-oxo-dGTP or dPTP alongside standard dNTPs. These analogs increase mispairing, particularly favoring transversions.
- Optimize Magnesium Concentration: Increase MgCl₂ concentration (e.g., to 7-8 mM) to reduce polymerase fidelity and increase overall mutation rate, which can help balance the spectrum.
- Switch Polymerases: Use a mutator strain polymerase like Mutazyme II, which is engineered for a more balanced mutational spectrum.
FAQ 2: Managing Mutation Hotspots
Q: Sequencing reveals recurrent mutations at specific positions (hotspots), reducing the diversity of my variant library. How can I distribute mutations more evenly? A: Hotspots often arise from sequence context dependence. Troubleshoot using these steps:
- Validate with a Control Sequence: Perform error-prone PCR on a standard, non-target control gene (e.g., lacZα) to determine if hotspots are inherent to your template or a result of the reaction conditions.
- Alter Sequence Context: If possible, redesign primers to slightly shift the amplified region or perform gene shuffling before mutagenesis to break up problematic motifs.
- Titrate Manganese: In Mn²⁺-based protocols, carefully titrate MnCl₂ (0.1-0.5 mM) as its concentration directly influences hotspot formation. Lower concentrations may reduce bias.
- Combine Methods: Use error-prone PCR for initial diversification, followed by a low-fidelity staggered extension process (StEP) or DNA shuffling to redistribute mutations.
Q: How can I precisely tune the number of mutations per gene/kb in my error-prone PCR library? A: Mutation frequency is controllable through reaction chemistry. Follow this protocol:
Protocol: Titration of Mutation Rate via Nucleotide Imbalance
- Prepare Base Reaction Mix (50 µL):
- 1X Thermostable polymerase buffer (Mg-free)
- 200 µM each dGTP and dATP
- Variable concentrations of dCTP and dTTP (see Table 1)
- 0.5 mM MnCl₂ (add last, from a fresh 10 mM stock)
- 1 ng/µL template DNA
- 0.3 µM each forward and reverse primer
- 1 U/µL Taq DNA Polymerase
- Thermocycling: 95°C for 2 min; [95°C for 30 sec, 55°C for 30 sec, 72°C for 1 min/kb] for 25-30 cycles; 72°C for 5 min.
- Clone & Sequence: Purify the product, clone into an appropriate vector, and sequence 10-20 random clones to calculate the average mutations/kb.
FAQ 4: Correcting for Sequence Context Dependence
Q: My data shows mutations are not random and depend heavily on local nucleotide sequence. How do I account for this in my experimental design and data analysis? A: Sequence context is an inherent biological bias. Mitigation strategies include:
- In Silico Modeling: Use tools like MutaGene or sequence-based mutation probability matrices to predict hotspots in your template before the experiment.
- Empirical Determination: Perform a small-scale pilot mutagenesis (as in FAQ 3) on your exact template to map its intrinsic mutation profile. Use this data to inform library size requirements.
- Post-PCR Normalization: For critical applications like drug target evolution, use high-throughput sequencing to characterize your initial library. You can then computationally down-sample over-represented variants or physically enrich under-represented ones via hybridization-based capture.
Table 1: Effect of dNTP Imbalance on Mutation Spectrum
| Condition | dCTP/dTTP (µM) | dATP/dGTP (µM) | Avg. Mutations/kb | Tv:Ts Ratio | Notes |
|---|---|---|---|---|---|
| Balanced | 200 | 200 | 2-4 | ~1.5:1 | Baseline, Ts slightly favored |
| Deplete dCTP/dTTP | 20 | 200 | 5-8 | ~0.8:1 | Increases A→G, G→A Transitions |
| Deplete dATP/dGTP | 200 | 20 | 5-8 | ~2.5:1 | Increases A→T, G→T Transversions |
| With 8-oxo-dGTP* | 200 | 150 (+50 8-oxo) | 10-15 | ~3:1 | Significantly boosts G→T transversions |
*Added as a partial substitute for dGTP.
Table 2: Common Mutational Hotspot Motifs in Error-Prone PCR
| Sequence Motif | Common Mutation(s) | Likely Cause | Mitigation Strategy |
|---|---|---|---|
| 5'-RGYA-3' (R=purine, Y=pyrimidine) | A→G, G→A | Polymerase slippage & misincorporation bias | Lower Mn²⁺, use mutator polymerase blend |
| Continuous GC-rich regions | C→T, G→A | Incomplete denaturation & stable mispairs | Add DMSO (3-5%), increase denaturation temp/time |
| 5'-TGG-3' / 5'-CCA-3' | T→C, C→T | Oxidation-prone guanine in triplet context | Use antioxidants in reaction, perform under inert gas |
Visualizations
Diagram: Error-Prone PCR Workflow & Bias Introduction
Diagram: Mutation Types - Transitions vs. Transversions
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Error-Prone PCR Library Construction
| Reagent | Function & Rationale | Example Product/Note |
|---|---|---|
| Mutagenic Polymerase Blend | Engineered or natural low-fidelity polymerases to introduce base misincorporations. | Mutazyme II (Agilent), GeneMorph II (Takara). Prefer blends over single Taq. |
| Manganese Chloride (MnCl₂) | Critical cofactor that reduces polymerase fidelity by promoting mispairing. Must be titrated precisely. | Prepare fresh 10-50 mM stock solutions in nuclease-free water. |
| Unbalanced dNTP Set | Individual dNTP solutions at high concentration (100 mM) to create precise nucleotide imbalances. | Use PCR-grade, pH-balanced stocks for reproducibility. |
| Nucleotide Analogues (8-oxo-dGTP, dPTP) | Directly promote specific mispairing events (e.g., 8-oxo-dGTP pairs with A, causing G→T transversions). | Use at 10-20% of total corresponding dNTP concentration. |
| High-Efficiency Cloning Kit | Essential for capturing the diverse, potentially damaged PCR library into E. coli with high efficiency. | NEBuilder HiFi DNA Assembly or Zero Blunt TOPO kits. Avoid low-efficiency ligation. |
| Next-Generation Sequencing Service/Kits | Required for accurate, quantitative analysis of the final mutation spectrum and library diversity. | Illumina MiSeq 300-cycle kits for amplicon sequencing of the library pool. |
Troubleshooting Guides & FAQs
Q1: During error-prone PCR (epPCR) library construction, my final DNA yield is consistently low. What could be the cause? A: Low yield in epPCR is often due to suboptimal magnesium (Mg2+) concentration or excessive mutation rate. Mg2+ is a cofactor for Taq polymerase; too little reduces efficiency. Conversely, a very high error rate (e.g., from excessive Mn2+) can introduce stop codons or destabilizing mutations, preventing full-length amplification.
- Troubleshooting Steps:
- Titrate Mg2+ and Mn2+: Perform a matrix experiment with Mg2+ (1-7 mM) and Mn2+ (0-0.5 mM).
- Check Template Quality: Ensure starting template is pure and quantitated accurately.
- Cycle Optimization: Reduce the number of PCR cycles to minimize accumulation of truncated products.
- Verify Polymerase: Use a polymerase validated for epPCR (e.g., Mutazyme II).
Q2: My NGS data shows a dramatic skew in sequence representation (some variants are overrepresented). How do I diagnose and correct this? A: Skew indicates bias introduced during PCR or transformation. Primary culprits are uneven PCR amplification and variance in E. coli transformation efficiency.
- Troubleshooting Steps:
- Limit PCR Cycles: Keep epPCR cycles to the minimum required for sufficient yield (often 25-30 cycles).
- Use High-Efficiency, Electrocompetent Cells: Ensure uniform transformation. Chemical competence can be highly variable.
- Normalize Post-Ligation: If constructing a ligation-based library, perform a gel extraction or use a size-selection bead clean-up to isolate correctly sized inserts before transformation.
- Analyze Early Timepoints: Sequence the plasmid library before transformation to distinguish PCR bias from transformation bias.
Q3: How can I tell if my mutagenesis library has sufficient diversity, or if it's too biased to be useful? A: You must calculate quantitative metrics from your NGS data. See the table of Key Metrics below for formulas and targets.
Key Metrics for Library Assessment (Quantitative Data)
| Metric | Formula / Description | Ideal Target | Indicates Problem If... |
|---|---|---|---|
| Library Size (Diversity) | # of unique transformants (colonies) | >10x intended screen capacity | <100x screen capacity; risk of missing hits. |
| Mutation Frequency | (Total # of mutations) / (Total # of bases sequenced) | 1-5 amino acid changes per gene | >10 AA changes; high fraction of non-functional clones. |
| Coverage (Depth) | Average # of reads per unique position | >50x read depth per variant | <10x; statistical confidence in variant calling is low. |
| Shannon Entropy (H) | H = -Σ (pi * log2(pi)); where p_i is frequency of variant i | Higher is better (H > 12 for large libs) | Low value; library is dominated by few sequences (high bias). |
| Gini Coefficient | Measures inequality in variant frequencies (0=perfect equality, 1=perfect inequality) | Closer to 0 (<0.3) | >0.5; severe frequency inequality, high bias. |
| % Wild-Type Sequence | (# of reads with 0 mutations) / (Total reads) | <5% for aggressive mutagenesis | >30%; mutagenesis protocol was inefficient. |
Experimental Protocol: Assessing Library Quality via NGS
Objective: To quantitatively measure bias, diversity, and mutation distribution in an epPCR library.
Materials:
- Purified epPCR plasmid library (post-transformation pool or pre-transformation amplicon).
- Primers for amplifying the variable region for NGS.
- High-fidelity PCR mix (e.g., Q5).
- NGS library prep kit (Illumina-compatible).
- Bioanalyzer/TapeStation.
- NGS platform (MiSeq recommended for depth).
Methodology:
- Library Amplification for NGS: Perform high-fidelity PCR (≤18 cycles) on your epPCR library using barcoded primers to add Illumina adapters and unique dual indices.
- Size Selection & Purification: Clean the PCR product with magnetic beads (e.g., SPRIselect) to remove primers and select the correct insert size. Verify fragment size on a Bioanalyzer.
- Quantitation & Pooling: Quantify the library by qPCR (for molarity) and pool with other libraries if applicable.
- Sequencing: Run on an Illumina MiSeq or NovaSeq with paired-end reads of sufficient length to cover the entire variable region. Aim for >100x average coverage.
- Bioinformatic Analysis:
- Demultiplex & Merge Reads: Use tools like
bcl2fastqandFLASH/PEAR. - Align to Reference: Align reads to the wild-type sequence using
BWAorBowtie2. - Variant Calling: Use
PoPoolation2or a custom script (e.g., inPythonwithpysam) to identify mutations per read. - Calculate Metrics: Compute Shannon Entropy, Gini Coefficient, mutation frequency, and coverage from the alignment file.
- Demultiplex & Merge Reads: Use tools like
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in epPCR/Diversity Assessment |
|---|---|
| Mutazyme II (or similar) | Engineered polymerase blend optimized for introducing random mutations with reduced bias. |
| MnCl2 Solution | Manganese ions induce misincorporation by Taq polymerase, driving the mutation rate. |
| Ultra-High Efficiency Electrocompetent E. coli | Maximizes transformation efficiency (>10^9 cfu/µg) to capture library diversity without bottlenecking. |
| Next-Generation Sequencing Kit (Illumina) | For deep sequencing the library to quantify bias and diversity metrics. |
| SPRIselect Magnetic Beads | For consistent, high-recovery size selection and purification of NGS libraries. |
| Q5 High-Fidelity DNA Polymerase | For accurate amplification of libraries for NGS prep without introducing additional mutations. |
Diagrams
Title: epPCR Library Construction and NGS Analysis Workflow
Title: Hierarchy of Key Bias and Quality Assessment Metrics
Engineering Diversity: Practical Protocols to Control and Direct Mutagenic Outcomes
Troubleshooting & FAQs for Error-Prone PCR Systems
Q1: My mutation rate is consistently lower than the kit's specification. What could be wrong? A: Low mutation frequency is common. First, verify template quality and quantity; excess template dilutes mutations. Ensure you are using the correct dNTP ratios if the kit supplies separate nucleotides for bias adjustment. A critical step is to strictly adhere to the recommended number of PCR cycles—exceeding cycles can lead to "jackpot" effects and wild-type carryover. Confirm the fidelity of your polymerase is appropriate; some systems use specialized low-fidelity polymerases (e.g., Mutazyme II, Taq) that are integral to the kit. Finally, run a post-PCR gel to check for nonspecific amplification, which can consume reagents.
Q2: I observe a strong sequence context bias, where mutations cluster in specific regions. How can I mitigate this? A: This addresses core thesis concerns of mutagenesis bias. Commercial kits attempt to minimize but not eliminate sequence-dependent bias. To combat this: 1) Use a kit that employs multiple mutagenesis enzymes (e.g., a blend of Mutazyme I & II) for more balanced mutation spectra. 2) Consider performing staggered or parallel reactions with different metal cofactors (e.g., Mn2+ concentration) and pooling the products. 3) For critical projects, a low-bias random mutagenesis kit that uses nucleotide analogues (e.g., dPTP) may be superior, though more expensive. This is a key factor in kit selection for unbiased library generation.
Q3: The final library diversity is insufficient for screening. What are the main culprits? A: Insufficient diversity often stems from bottlenecks. Calculate and minimize the number of E. coli transformants; you must achieve at least 3-5x library size coverage. Use high-efficiency electrocompetent cells. During PCR, ensure the input DNA is minimal (as low as 10-100 ng) to favor mutated strands. A critical troubleshooting step is to sequence 20-50 individual clones from the library to empirically determine mutation rate and distribution before large-scale transformation.
Q4: How do I choose between a kit-based system and a custom protocol? A: Kits (e.g., from Agilent, Thermo Fisher, Jena Bioscience) offer reproducibility, convenience, and optimized buffers, which is vital for standardized research. Custom protocols (using Mn2+, unbalanced dNTPs) allow fine-tuning for specific bias reduction but require extensive optimization. For most drug development professionals, a commercial kit with a validated, consistent error rate is preferable for scalable, reproducible work. Your choice should be guided by the thesis need to control versus characterize bias.
Comparative Data Table: Key Commercial Error-Prone PCR Systems
| Kit/System Name (Supplier) | Core Mutagenesis Method | Theoretical Mutation Rate (Range) | Key Advantage | Noted Bias Concerns |
|---|---|---|---|---|
| GeneMorph II Random Mutagenesis Kit (Agilent) | Mutazyme II enzyme blend | 0-16 mutations/kb (adjusted by template amount) | Tunable rate via input DNA; high-fidelity backbone. | AT bias reported in some sequence contexts. |
| Diversify PCR Random Mutagenesis Kit (Takara Bio) | Modified Taq polymerase + uneven dNTPs | 2-8 mutations/kb | Simple, cost-effective; good for low-to-medium rates. | Standard Taq bias (preference for transitions). |
| JBS Random Mutagenesis Kit (Jena Bioscience) | Mutazyme I & II, and nucleotide analogues | 1-40 mutations/kb | Very broad rate range; multiple enzyme options. | Kit selection crucial to match enzyme to desired bias. |
| NXGen Random Mutagenesis Kit (Lucigen) | Proprietary mutagenic polymerase | 1-10 mutations/kb | Designed for low sequence context bias. | Newer system; independent validation data is growing. |
| Thermo Scientific GeneArt Mutagenesis Kit | Not publicly detailed (proprietary enzyme mix) | Adjustable | Integrated system for library construction & cloning. | Proprietary nature makes bias mechanism less transparent. |
Standardized Protocol: Error-Prone PCR with a Commercial Kit
This protocol is generalized for kits like the Agilent GeneMorph II.
1. Reaction Setup:
- Template: Dilute plasmid or linear DNA to 10 ng - 1 µg in nuclease-free water. Use the lower end for higher mutation rates.
- Master Mix: On ice, combine:
- 5 µL 10x Mutazyme II Reaction Buffer
- 1 µL (e.g., 10 ng) template DNA
- 1 µL 40 mM dNTP Mix (supplied)
- 1 µL Primer 1 (10 µM)
- 1 µL Primer 2 (10 µM)
- 1 µL Mutazyme II DNA Polymerase (2.5 U/µL)
- 39 µL Nuclease-Free Water
- Total Volume: 50 µL.
2. Thermal Cycling:
- Initial Denaturation: 95°C for 2 min.
- Cycling (25-30 cycles):
- Denature: 95°C for 30 sec.
- Anneal: 55-65°C (primer-specific) for 30 sec.
- Extend: 72°C for 1 min/kb.
- Final Extension: 72°C for 10 min.
- Hold: 4°C.
3. Post-PCR Processing:
- Run 5 µL on agarose gel to confirm amplification.
- Purify the PCR product using a spin column-based PCR purification kit.
- Proceed to downstream cloning (digestion, ligation) into your expression vector.
Visualization: Error-Prone PCR Experimental Workflow
Title: Error-Prone PCR Library Construction Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Error-Prone PCR |
|---|---|
| Low-Fidelity Polymerase Blend (e.g., Mutazyme II) | Engineered enzyme with reduced proofreading to base misincorporation, the core mutagenic driver. |
| Unbalanced dNTP Stocks (e.g., high dATP, dTTP) | Increases probability of misincorporation by unbalancing natural nucleotide ratios. |
| Manganese Chloride (Mn2+) Solution | Often used in custom protocols; Mn2+ promotes nucleotide misincorporation by reducing polymerase fidelity. |
| High-Efficiency Electrocompetent Cells (>10^9 cfu/µg) | Essential for capturing the full diversity of the mutant library without bottlenecking. |
| PCR Purification Kit (Spin Column) | Removes excess primers, dNTPs, and enzymes post-amplification before cloning steps. |
| Broad-Host-Range Cloning Vector | Allows expression of the mutated gene in the desired screening host (e.g., E. coli, yeast). |
| Next-Generation Sequencing (NGS) Service/Kits | For comprehensive analysis of final library mutation rate, spectrum, and bias. |
Troubleshooting Guides & FAQs
FAQ: General Principles
Q: What is the fundamental relationship between Mn2+, Mg2+, and mutation rate in error-prone PCR? A: Mn2+ is the primary mutagenic driver. It promotes misincorporation by reducing the fidelity of polymerase, as Mn2+ ions can be mistakenly incorporated by the polymerase instead of Mg2+. Mg2+ is essential for standard polymerase activity and fidelity. The ratio of Mn2+:Mg2+, along with absolute concentrations, critically determines the mutation frequency and spectrum. High dNTP concentrations can further increase the error rate by reducing polymerase selectivity.
Q: What is a typical starting point for Mn2+ concentration to achieve a moderate mutation rate? A: A common starting concentration is 0.1-0.5 mM MnCl2, with a standard 1.5 mM MgCl2. This often yields a mutation rate in the range of 0.1-1% per base. Optimization is required for each system.
Q: How do I adjust my protocol to bias mutations toward transitions (AG, CT) vs. transversions? A: Increasing the concentration of Mn2+ relative to Mg2+ favors transversions. For a bias toward transitions, use a lower Mn2+:Mg2+ ratio and consider supplementing with nucleotide analogs like 8-oxo-dGTP or dPTP, though this moves beyond simple ion manipulation.
Troubleshooting Guide
Problem: No PCR product or drastically reduced yield after adding Mn2+.
- Cause: Excessive Mn2+ concentration is inhibiting polymerase activity.
- Solution: Titrate MnCl2 in 0.05 mM increments from 0.05 mM to 0.8 mM. Keep MgCl2 constant at 1.0-1.5 mM initially. Ensure the total divalent cation (Mn2+ + Mg2+) concentration is within the polymerase's functional range (typically 1-4 mM).
Problem: Mutation rate is too low despite adding Mn2+.
- Cause 1: Mg2+ concentration is too high, outcompeting Mn2+.
- Solution: Gradually reduce MgCl2 concentration (e.g., from 2.0 mM to 0.7 mM) while maintaining a constant Mn2+ level (e.g., 0.3 mM). Monitor yield carefully.
- Cause 2: dNTP concentration is too low.
- Solution: Increase dNTP concentration (e.g., from 200 µM to 600 µM each). Higher dNTPs can reduce fidelity.
Problem: Mutation rate is too high, generating excessive stop codons and non-functional variants.
- Cause 1: Mn2+ concentration is too high.
- Solution: Decrease MnCl2 concentration.
- Cause 2: dNTP concentration is excessively high.
- Solution: Reduce dNTP concentration to standard levels (e.g., 200 µM each).
- General Adjustment: Increase the Mg2+:Mn2+ ratio to favor fidelity over mutagenesis.
Problem: Mutational bias is skewed, not producing the desired diversity.
- Cause: The Mn2+/Mg2+/dNTP balance creates a specific misincorporation profile.
- Solution: Systematically vary the ratios. See the table below for targeted adjustments. Consider using specialized polymerases with inherent mutational biases (e.g., Mutazyme II) if ion tuning is insufficient.
Data Presentation
Table 1: Effect of Reaction Component Titration on Mutation Rate and Spectrum
| Component | Typical Concentration Range | Effect on Mutation Rate (Per Base Pair) | Influence on Mutation Spectrum | Key Consideration |
|---|---|---|---|---|
| MnCl2 | 0.05 - 0.8 mM | Primary driver. Increases linearly then plateaus. | Higher [Mn2+] favors transversions (A:T→C:G; G:C→T:A). | Inhibits polymerase >1.0 mM; requires Mg2+ co-presence. |
| MgCl2 | 0.5 - 4.0 mM | Inverse relationship with fidelity. Low [Mg2+] increases error rate. | Lower [Mg2+] can increase transition frequency. | Absolute requirement for polymerase activity. Optimize with Mn2+. |
| dNTPs (each) | 0.1 - 1.0 mM | Moderate increase with higher concentration. | Can alter bias based on relative concentrations (e.g., high dGTP→C→T). | Excess dNTPs chelate Mg2+/Mn2+, affecting availability. |
| Mg2+:Mn2+ Ratio | 1:1 to 20:1 | Critical control parameter. Lower ratio = higher rate. | Lower ratio increases transversion frequency. | The most important strategic variable for fine-tuning. |
Table 2: Example Optimization Matrix for Targeted Mutation Rates
| Target Mutation Rate | Suggested [Mn2+] | Suggested [Mg2+] | Suggested [dNTPs] each | Expected Outcome |
|---|---|---|---|---|
| Low (0.1-0.3%) | 0.05 - 0.1 mM | 1.5 - 2.0 mM | 200 µM | Maintains library functionality; low diversity. |
| Medium (0.5-1.0%) | 0.2 - 0.4 mM | 1.0 - 1.5 mM | 300 - 400 µM | Balanced diversity and functional protein yield. |
| High (2-4%) | 0.5 - 0.7 mM | 0.7 - 1.0 mM | 500 - 600 µM | High diversity, but significant fraction of non-functional variants. |
Experimental Protocols
Protocol 1: Standard Error-Prone PCR with Mn2+ Titration
Objective: Generate a library with a gradient of mutation frequencies.
- Master Mix (per 50 µL rxn): 1X Polymerase Buffer (Mg-free), 0.2 mM each dNTP, 0.3 µM each primer, 10-50 ng template, 2.5 U Taq polymerase.
- Variable Component: Prepare 5 tubes. Add MnCl2 from a 10 mM stock to final concentrations of 0, 0.1, 0.3, 0.5, and 0.7 mM.
- Constant Component: To all tubes, add MgCl2 to a final concentration of 1.5 mM.
- PCR Cycling: Initial denaturation: 95°C, 2 min. Then 30 cycles of: 95°C for 30 sec, 55°C for 30 sec, 72°C for 1 min/kb. Final extension: 72°C, 5 min.
- Analysis: Purify products and sequence 5-10 clones from each condition to calculate mutation rate.
Protocol 2: Fine-Tuning Mutational Bias by Adjusting Mg2+:Mn2+ Ratio
Objective: Shift the mutation spectrum toward more transitions.
- Setup: Use a fixed, moderate MnCl2 concentration (e.g., 0.3 mM).
- Titration: Set up reactions with MgCl2 at 0.7, 1.1, 1.5, and 2.0 mM. This creates Mg2+:Mn2+ ratios of ~2.3, 3.7, 5, and 6.7.
- Constant: Keep dNTPs at 200 µM each.
- PCR: Perform amplification as in Protocol 1.
- Analysis: Deep sequence the resulting libraries (>10,000 reads) to analyze the relative proportions of transition vs. transversion mutations.
Mandatory Visualization
Optimizing Error-Prone PCR Protocol
Mutagenesis Mechanism of Mn2+ in PCR
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Error-Prone PCR | Key Consideration |
|---|---|---|
| Manganese Chloride (MnCl2) | Primary mutagenic agent. Causes polymerase misincorporation by distorting active site geometry. | Use a fresh, aqueous stock. Concentration is critical; titrate carefully. |
| Magnesium Chloride (MgCl2) | Essential cofactor for polymerase activity. Competes with Mn2+ to modulate fidelity and yield. | Optimize concentration relative to Mn2+. Often supplied in buffer; may need Mg-free buffer. |
| dNTP Mix (high concentration) | Substrates for DNA synthesis. Elevated concentrations can reduce polymerase selectivity. | High dNTPs chelate divalent cations; adjust Mg2+/Mn2+ accordingly. |
| Low-Fidelity Polymerase (e.g., Taq) | Naturally lower fidelity than high-fidelity enzymes; more responsive to Mn2+ manipulation. | Avoid proofreading polymerases (e.g., Pfu) as they correct errors. |
| Mg2+/Mn2+-Free Reaction Buffer | Allows precise, independent control over divalent cation concentrations. | Essential for systematic optimization. |
| Nucleotide Analogs (e.g., 8-oxo-dGTP) | Can be used in conjunction with Mn2+ to create specific mutational biases (e.g., A→C). | Expands mutational spectrum beyond what ions alone achieve. |
| PCR Purification Kit | To clean up product before downstream cloning or sequencing. | Removes excess salts, enzymes, and nucleotides that could interfere. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My error-prone PCR (epPCR) with a mutant Taq polymerase (e.g., Mutazyme II) is yielding no detectable product. What are the primary causes? A: This is commonly due to incorrect Mg2+ concentration or an incompatible dNTP bias mixture.
- Actionable Steps:
- Titrate Mg2+: Perform a MgCl2 gradient from 1.5 mM to 7 mM. Engineered polymerases often require higher, non-standard concentrations.
- Verify dNTP Ratios: Confirm the preparation of your biased dNTP pool. A common starting point is 0.2 mM each dNTP, but for increased mutagenesis, use uneven ratios (e.g., 1 mM dATP, 0.2 mM each of dCTP, dGTP, dTTP).
- Check Template Quality: Ensure template DNA is pure and at an optimal concentration (10-100 ng for a 1-3 kb plasmid).
Q2: The mutation rate from my epPCR is significantly lower than the enzyme's specification. How can I increase it? A: The mutation rate is a function of polymerase error rate and reaction conditions.
- Actionable Steps:
- Increase Manganese: Add MnCl2 (0.1-0.5 mM) to the reaction. Mn2+ is a known mutagenic cofactor that promotes misincorporation. Note: This can reduce yield.
- Adjust dNTP Ratios Further: Implement a more aggressive bias (e.g., 1 mM dATP/dTTP, 0.1 mM dCTP/dGTP).
- Cycle Optimization: Increase the number of PCR cycles (e.g., 30-40 cycles).
Q3: I am using a specialist enzyme (e.g., KAPA HiFi) for PCR of GC-rich regions prior to mutagenesis, but specificity is poor. A: GC-rich amplification requires additives and specific thermal cycling parameters.
- Actionable Steps:
- Add Enhancers: Include DMSO (3-10%), Betaine (1-1.5 M), or GC-rich specific buffers supplied by the manufacturer.
- Use a Two-Step PCR: Combine a high-fidelity enzyme for template amplification with a mutator polymerase in a subsequent, separate epPCR step to avoid bias in the initial product.
- Optimize Annealing: Perform a gradient PCR to find the optimal annealing temperature, which is often higher for GC-rich templates.
Q4: How do I choose between different commercial mutant Taq polymerases for my directed evolution project? A: Selection depends on the desired mutation spectrum and rate, as outlined in the table below.
Table 1: Comparison of Engineered Polymerases for Error-Prone PCR
| Polymerase (Example) | Key Mutation Bias | Avg. Error Rate (mutations/kb) | Optimal [Mg2+] | Recommended [Mn2+] | Best For |
|---|---|---|---|---|---|
| Mutazyme II | AT → GC & GC → AT | 16 - 40 | 7 mM | 0 - 0.5 mM | Balanced spectrum, high-rate mutagenesis |
| Genemorph II | AT → TA & GC → CG | 2 - 16 | 2 - 4 mM | Not required | Lower mutation rates, transversion bias |
| Taq Pol (wild-type) | AT → GC | ~1.1 | 1.5 - 2.5 mM | Not standard | Baseline, low-fidelity applications |
| KAPA HiFi | Minimal (High-Fid.) | ~0.03 | 1.5 - 2.5 mM | Never | High-fidelity template amplification |
Experimental Protocols
Protocol 1: Standard Error-Prone PCR using a Mutant Taq Polymerase
- Objective: Generate a library of mutated DNA fragments.
- Reagents: Mutant Taq polymerase (e.g., Mutazyme II), 10X Mutagenic Buffer, biased dNTP mix, template DNA, primers.
- Method:
- Prepare a 50 µL reaction: 5 µL 10X Mutagenic Buffer, 1 µL biased dNTP mix (e.g., 10 mM total), 10-100 ng template, 10 pmol each primer, 1.0 µL Mutazyme II, adjust to 50 µL with nuclease-free water.
- Thermal Cycling: Initial denaturation: 95°C for 2 min. Then 30 cycles of: 95°C for 30 sec, 55-65°C (primer-specific) for 30 sec, 72°C for 1 min/kb. Final extension: 72°C for 5 min.
- Purify the PCR product using a spin column before downstream cloning.
Protocol 2: Two-Step PCR for GC-Rich Template Mutagenesis
- Objective: Amplify a difficult template with high fidelity, then introduce mutations in a second, separate reaction.
- Reagents: High-fidelity polymerase (e.g., KAPA HiFi), epPCR polymerase, two separate buffer systems.
- Method:
- Step 1 (High-Fidelity PCR): Amplify the GC-rich target using KAPA HiFi per manufacturer's instructions, including 3% DMSO. Purify the amplicon.
- Step 2 (epPCR): Use 10-50 ng of the purified Step 1 product as template in a standard epPCR reaction (as in Protocol 1). This prevents the wild-type sequence bias from dominating the final library.
Visualizations
Title: Enzyme Selection & Experimental Workflow for Tailored Mutagenesis
Title: Mechanisms of Engineered Polymerases for Bias Control
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Tailored Mutagenesis Experiments
| Reagent / Material | Function & Rationale |
|---|---|
| Mutant Taq Polymerase Kit (e.g., Mutazyme II, Genemorph II) | Core enzyme with inherent high error rate and specific mutational bias for generating diversity. |
| High-Fidelity Polymerase (e.g., KAPA HiFi, Q5) | For faithful amplification of template DNA, especially critical for GC-rich targets, before mutagenesis. |
| MgCl2 Solution (25-100 mM) | Essential cofactor. Concentration must be optimized for each mutant polymerase, often above standard levels. |
| MnCl2 Solution (5-10 mM) | Mutagenic cofactor that stabilizes non-complementary base pairing, used to further increase error rate. |
| Biased dNTP Mixtures | Pre-mixed or custom dNTP solutions with unequal concentrations to steer the mutation spectrum. |
| PCR Additives (DMSO, Betaine) | Reduce secondary structure in DNA, crucial for amplifying high-GC templates prior to mutagenesis. |
| PCR Purification Kit | For clean-up of epPCR products to remove enzymes, salts, and primers before downstream cloning steps. |
| Mutagenic Buffer (10X) | Proprietary buffer supplied with enzyme, often containing optimized salt concentrations for mutagenesis. |
Technical Support Center: Troubleshooting & FAQs
Q1: During error-prone PCR (epPCR) with nucleotide analogs like 8-oxo-dGTP and dPTP, my product yield is extremely low or nonexistent. What are the primary causes and solutions? A: Low yield is commonly due to polymerase stalling or inability to incorporate the analog.
- Cause 1: Polymerase Incompatibility. Standard Taq polymerase may inefficiently incorporate bulkier analogs.
- Solution: Use engineered or specialized polymerases (e.g., Mutazyme II, Therminator IX) with broader substrate tolerance. See Protocol 1.
- Cause 2: Excessive Analog Concentration. High levels can completely inhibit extension.
- Solution: Titrate the analog. Start with a molar ratio of 1:10 (analog:dNTP) and optimize. See Table 1 for typical working concentrations.
- Cause 3: Incorrect Mg²⁺ Concentration. Mg²⁺ is a crucial cofactor, and analogs may alter optimal conditions.
- Solution: Optimize Mg²⁺ concentration in the range of 2–8 mM.
Q2: My sequencing data shows a very low mutation frequency despite using analogs. How can I increase the mutagenesis rate? A: This indicates insufficient analog incorporation.
- Cause 1: Analog Ratio Too Low. The native dNTPs are outcompeting the analogs.
- Solution: Gradually increase the proportion of the problem analog in the nucleotide mix. Do not exceed thresholds that cause primer extension to fail (see Table 1).
- Cause 2: PCR Cycle Number Too Low. With lower incorporation efficiency, more cycles are needed to accumulate mutations.
- Solution: Increase PCR cycles to 35–40, ensuring sufficient template for later cycles.
- Cause 3: Analog Degradation. Some analogs (e.g., dPTP) are less stable.
- Solution: Prepare analog stocks fresh from powder, aliquot, and store at -80°C. Avoid multiple freeze-thaw cycles.
Q3: I observe a strong mutational bias (e.g., only transitions) instead of the promised transversion diversity from an analog like dPTP. Why? A: This directly relates to the thesis context of addressing mutagenesis bias. Bias occurs due to persistent polymerase preference and analog mispairing rules.
- Cause 1: Polymerase Fidelity and Proofreading. Even permissive polymerases have inherent preferences. Proofreading polymerases will excise mismatched analogs.
- Solution: Use non-proofreading, mutagenesis-optimized polymerases. Combine multiple analogs at sub-saturating levels to diversify mismatch patterns. See Protocol 2.
- Cause 2: Incomplete Native dNTP Exhaustion. If the native dNTP corresponding to the analog is still abundant, it will be incorporated preferentially.
- Solution: For a targeted approach, reduce or omit the competing native dNTP (e.g., omit dGTP when using 8-oxo-dGTP). Monitor yield closely.
Q4: How do I calculate and verify the mutation frequency and spectrum from my epPCR experiment with analogs? A: This requires sequencing and analysis.
- Solution: Clone the epPCR product and sequence 20-50 individual clones. Calculate mutation frequency as (total mutations / total base pairs sequenced). Categorize mutations as transitions (Ts) or transversions (Tv) and plot the spectrum.
- Protocol 3: Sanger Sequencing Analysis of Mutation Spectrum.
- Ligate epPCR product into a cloning vector and transform.
- Pick 30-50 colonies for colony PCR and Sanger sequencing.
- Align sequences to the original template using software (e.g., Geneious, SnapGene).
- Tally all point mutations. Exclude the vector region.
- Calculate: Mutation Frequency = (Total Mutations) / (Template Length in bp x Number of Clones Sequenced).
- Categorize each mutation as A:T→G:C, G:C→A:T (Transitions), or as one of the four Transversion types.
- Protocol 3: Sanger Sequencing Analysis of Mutation Spectrum.
Data Presentation
Table 1: Common Nucleotide Analogs for epPCR and Optimization Guide
| Analog | Target Mutation(s) | Typical Molar Ratio (Analog:dNTP) | Expected Mutation Frequency Range (%) | Key Consideration |
|---|---|---|---|---|
| 8-oxo-dGTP | Primarily G:C→T:A Transversions | 1:5 to 1:10 | 0.5 - 2.0 | Also induces some G→A transitions; can be combined with dPTP. |
| dPTP (2'-Deoxy-P-nucleoside-5'-Triphosphate) | A:T→G:C & G:C→A:T Transitions | 1:3 to 1:5 | 0.7 - 3.0 | Pair with non-proofreading polymerase. Unstable, use fresh. |
| 5-Bromo-dUTP | A:T→G:C Transitions | 1:5 to 1:20 | 0.2 - 1.5 | Mimics T; mispairs with G. UV-sensitive. |
| 6-(2-Deoxy-β-D-ribofuranosyl)-3,4-dihydro-8H-pyrimido[4,5-c][1,2]oxazin-7-one triphosphate (dZ) | A:T→C:G & T:A→C:G Transversions | 1:10 to 1:30 | 0.1 - 0.8 | Novel analog for underrepresented transversions. Low incorporation efficiency. |
Experimental Protocols
Protocol 1: Standard epPCR with Nucleotide Analogs (50 µL Reaction)
- Objective: Introduce random mutations via polymerase incorporation of nucleotide analogs.
- Reagents: Template DNA (10-50 ng), mutagenic polymerase (1.25 U), 10x reaction buffer, dNTP mix (variable), nucleotide analog stock, forward and reverse primers (0.5 µM each), nuclease-free water.
- Steps:
- Prepare Master Mix on ice: Water (variable), 10x Buffer (5 µL), dNTP Mix (e.g., 200 µM each native dNTP), Nucleotide Analog (e.g., 40 µM 8-oxo-dGTP), Primers (0.5 µM final each), Polymerase (1.25 U).
- Add template DNA to individual tubes.
- Aliquot master mix into each tube. Mix gently.
- Thermocycling: Initial Denaturation: 95°C, 2 min. Then 35 cycles of: Denature (95°C, 30 sec), Anneal (Tm of primers, 30 sec), Extend (72°C, 1 min/kb). Final Extension: 72°C, 5 min. Hold at 4°C.
- Purify PCR product using a spin column kit.
Protocol 2: Bias-Minimized epPCR Using a Dual Analog System
- Objective: Broaden mutation spectrum by combining analogs targeting complementary bases.
- Reagents: As in Protocol 1, plus two compatible analogs (e.g., 8-oxo-dGTP and dPTP).
- Steps:
- Prepare a Balanced Nucleotide Mix: Reduce native dGTP and dATP to 100 µM. Add 8-oxo-dGTP to 20 µM and dPTP to 30 µM. Keep dCTP and dTTP at 200 µM.
- Set up reaction as in Protocol 1, using the balanced nucleotide mix.
- Use a highly processive, analog-tolerant polymerase (e.g., Mutazyme II).
- Reduce cycling to 30 cycles to minimize wild-type amplification bias.
- Clone and sequence to assess spectrum (see Protocol 3).
Mandatory Visualization
Diagram 1: Nucleotide Analog Integration Workflow in epPCR
Diagram 2: Mutational Outcomes from Common Analogs
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Mutazyme II DNA Polymerase | A proprietary blend of Taq and a mutagenic polymerase. Engineered for high processivity and broad nucleotide analog incorporation, reducing bias. |
| Therminator IX DNA Polymerase | A 9°N polymerase variant with low discrimination against modified nucleotides. Ideal for bulky analogs like dZ. |
| 8-oxo-dGTP Sodium Salt | Direct chemical mutagen. In its syn conformation, it pairs with A, leading to G→T transversions, expanding beyond transition-only bias. |
| dPTP (Na Salt) | A degenerate base analog that pairs with both A and G, primarily inducing A→G and G→A transitions to increase mutation load. |
| 5-Bromo-2’-deoxyuridine 5’-Triphosphate (BrdUTP) | Thymidine analog. The bromine atom shifts electron density, increasing the chance of mispairing with G during replication (A→G transitions). |
| Custom dNTP:Analog Balanced Mix | Pre-mixed solutions with optimized ratios of native dNTPs to analogs (e.g., 1:5 8-oxo-dGTP:dGTP) to ensure consistent mutagenic input across experiments. |
| High-Efficiency Cloning Kit | Essential for generating clonal populations from heterogeneous epPCR products for accurate sequence analysis of mutation spectrum. |
Technical Support Center: Troubleshooting & FAQs
Frequently Asked Questions
Q1: Our combined mutagenesis library shows drastically skewed variant representation. What are the primary causes and how can we correct this?
A1: Skewed libraries typically result from unequal incorporation efficiency or biased amplification. Implement the following corrective protocol:
- Quantify Oligonucleotide Input: Use UV spectrophotometry (NanoDrop) to ensure equimolar pooling of mutagenic oligonucleotides. Re-synthesize any oligo with significant deviation (>15% from mean concentration).
- Optimize PCR Conditions: For the primary epPCR step, reduce template concentration to 0.5 pM and use a proofreading polymerase with dNTPs supplemented with 0.1-0.5 mM MnCl₂. This helps balance mutation distribution.
- Post-Synthesis Purification: PAGE-purify all synthetic oligonucleotides to remove truncated sequences that cause preferential amplification.
- Library Analysis: Sequence 50-100 colonies via Sanger sequencing to calculate Shannon Entropy (H'). A well-balanced library typically has H' > 3.5 for a 1,000-member pool.
Q2: We observe poor splicing efficiency when joining epPCR fragments with oligonucleotide cassettes via Gibson Assembly. What troubleshooting steps are recommended?
A2: This is often due to suboptimal overlap design or reaction conditions.
- Verify Overlap Lengths: Ensure 20-25 bp homologous overlaps for Gibson Assembly. For Golden Gate assembly, ensure the Type IIS restriction site (e.g., BsaI) is absent from your insert and backbone.
- Optimize Fragment Ratios: Use a molar insert:vector ratio of 3:1. Prepare fragments as per the table below:
Fragment Optimal Length Range Recommended Purification epPCR product 200-800 bp Gel extraction & column clean-up Oligo cassette 40-100 bp PAGE purification Linearized vector 3-5 kb DpnI digestion + gel extraction - Incubation Time: Extend assembly incubation time to 60 minutes at 50°C for Gibson Assembly.
Q3: How do we minimize background (wild-type) carryover in the final mutant library?
A3: Background arises from incomplete digestion of template DNA or inefficient mutant strand selection.
- Template Digestion: Following epPCR, treat the product with DpnI (restricts methylated template DNA) for 3 hours using 1 U/µL enzyme. Verify digestion on an agarose gel.
- Strand Selection: If using oligonucleotide-mediated mutagenesis with a phosphorothioate-based selection, ensure β-mercaptoethanol is fresh and added to the cleavage reaction at 10 mM final concentration.
- Transformation Control: Always include a "ligation-only" negative control. Acceptable background is <5% wild-type colonies as determined by diagnostic restriction digest or sequencing.
Q4: What is the expected combined mutation rate, and how is it quantified?
A4: The combined method aims for 2-8 mutations per kb. Quantification protocol:
- Sequencing Sampling: Pick 20 random colonies from the final library for Sanger sequencing.
- Data Analysis: Align sequences to the wild-type gene. Calculate the average number of mutations per kb.
- Targeted vs. Random: Distinguish mutations from epPCR (random, mostly transitions) from those introduced by oligonucleotides (targeted, defined). See expected distribution table:
Mutation Source Typical Rate (mutations/kb) Primary Type Control Parameter Error-prone PCR 1-5 A/T → G/C transitions Mn²⁺ concentration, dNTP bias Oligonucleotide Cassette 1-3 (per oligo) Defined substitutions/insertions Oligo design, annealing temp Total Combined 2-8 Mixed See troubleshooting Q1
Q5: The transformation efficiency of our final assembled library is too low for adequate coverage. How can we improve yield?
A5: Low efficiency points to assembly toxicity or poor electrocompetent cells.
- Desalt Assembly Mix: Prior to transformation, desalt the Gibson/Golden Gate assembly mixture using a spin column or drop dialysis against sterile water.
- Electrocompetent Cells: Use high-efficiency E. coli cells (≥ 1 x 10⁹ cfu/µg). Thaw cells on ice completely and use pre-chilled cuvettes.
- Recovery: Increase recovery time to 1.5 hours at 37°C with shaking (900 µl SOC medium). Plate various volumes (1, 10, 100 µl) to ensure accurate counting.
- Calculate Library Coverage: Use the formula: Coverage = (Number of Colonies × Average Insert Size) / Library Diversity. Aim for a coverage factor >10x.
Experimental Protocols
Protocol 1: Two-Stage Combined Mutagenesis Workflow
Stage 1: Low-Bias Error-Prone PCR
- Reaction Mix (50 µL):
- 10-50 ng template plasmid (linearized)
- 0.3 µM each forward/reverse primer (flanking insertion site)
- 1X proprietary epPCR buffer (commercial kit, e.g., Genemorph II)
- 0.2 mM each dATP/dGTP, 1.0 mM each dCTP/dTTP (creates A/T bias)
- 0.1 mM MnCl₂
- 1.25 U mutagenic polymerase blend
- Thermocycling:
- 95°C for 2 min.
- 30 cycles of: 95°C for 30 sec, 55°C for 30 sec, 72°C for 1 min/kb.
- 72°C for 10 min.
- Clean-up: Purify PCR product with magnetic beads. Digest with DpnI (3h, 37°C) to remove template.
Stage 2: Oligonucleotide Cassette Splicing via Golden Gate Assembly
- Design: Order oligonucleotides as complementary pairs with 5' overhangs matching the BsaI-digested vector and internal mutated sequence.
- Annealing: Mix each oligo pair (100 µM each) in 10 mM Tris, 50 mM NaCl, 1 mM EDTA. Heat to 95°C for 2 min, cool slowly to 25°C (0.1°C/sec).
- Assembly Reaction (20 µL):
- 50 ng DpnI-treated epPCR fragment (as vector backbone)
- 0.1 pmol of each annealed duplex oligonucleotide cassette
- 1X T4 DNA Ligase Buffer
- 10 U BsaI-HFv2
- 400 U T4 DNA Ligase
- Cycling: 30 cycles of (37°C for 2 min, 16°C for 5 min), then 60°C for 10 min, 80°C for 20 min.
Protocol 2: Library Quality Assessment by NGS
- Amplify Library for Sequencing: Perform a high-fidelity PCR (10 cycles) using primers adding Illumina adapters and sample indexes.
- Purify & Quantify: Clean amplicons via dual-sided bead selection. Quantify by Qubit and qPCR (Kapa Library Quant Kit).
- Sequencing: Run on a MiSeq (2x250 bp) to achieve >100x coverage per variant.
- Bioinformatic Analysis:
- Demultiplex reads (bcl2fastq).
- Align to reference (BWA-MEM).
- Call variants (GATK HaplotypeCaller).
- Calculate mutation spectrum and library entropy using custom Python scripts (available on GitHub repository linked in thesis).
Visualizations
Diagram 1: Combined Mutagenesis Experimental Workflow
Diagram 2: Mutagenesis Bias Correction Strategy
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Combined Mutagenesis | Key Consideration |
|---|---|---|
| Mutazyme II DNA Polymerase | Engineered epPCR enzyme with broad mutational spectrum. | Reduces bias compared to Taq-based methods. Use provided buffer. |
| MnCl₂ Solution (10 mM) | Adds to PCR buffer to increase error rate by promoting misincorporation. | Critical for tuning mutation frequency. Titrate (0.1-0.5 mM final). |
| Biased dNTP Mixes | Pre-mixed solutions with unequal dNTP concentrations (e.g., high dCTP/dTTP). | Counters natural polymerase bias to achieve uniform mutation distribution. |
| Phosphorothioate-Modified Oligos | Oligonucleotides with sulfur-substituted backbone for strand selection (Nickase resistance). | Enriches for mutant strands. Requires specific cleavage with Iodine/EtOH. |
| BsaI-HFv2 Restriction Enzyme | High-fidelity Type IIS enzyme for Golden Gate assembly. Creates defined 4-bp overhangs. | Minimizes star activity. Essential for modular cassette assembly. |
| Gibson Assembly Master Mix | All-in-one mix of exonuclease, polymerase, and ligase for seamless assembly. | Best for larger fragments (>200 bp). Desalt before transformation. |
| Electrocompetent NEB 10-beta E. coli | High-efficiency cells for library transformation. | Essential for achieving >10⁹ library size. Must be kept at -80°C until use. |
| KAPA Library Quantification Kit | qPCR-based assay for accurate sequencing library quantification. | Ensures optimal cluster density on Illumina sequencers for QC. |
Technical Support Center
Troubleshooting Guides & FAQs
FAQ Category 1: PCR Optimization for Library Diversity
Q1: My error-prone PCR (epPCR) library shows very low mutational diversity. What are the key parameters to adjust?
- A: Low diversity often stems from an insufficient mutation rate. To increase it, systematically adjust your epPCR conditions. Critical variables include:
- MgCl₂ Concentration: Mg²⁺ is crucial for Taq polymerase fidelity. Increasing it (e.g., from 1.5 mM to 7 mM) can decrease fidelity.
- MnCl₂ Addition: Mn²⁺ induces misincorporation. Titrate from 0 to 0.5 mM.
- dNTP Imbalance: Unequal dNTP concentrations (e.g., increasing dATP and dTTP while decreasing dCTP and dGTP, or vice versa) bias misincorporation.
- Polymerase Choice: Use Taq polymerase (low fidelity) over high-fidelity enzymes.
- Template Amount: Use low template concentration (e.g., < 100 ng per 50 µL reaction) to reduce wild-type carryover.
- Protocol: Standardized epPCR Optimization Test
- Set up a matrix of 4 reactions (50 µL each):
- Reaction A (Baseline): 1x Buffer, 1.5 mM MgCl₂, 0.2 mM each dNTP, 0.3 µM primers, 50 ng template, 1.25 U Taq.
- Reaction B (High Mg²⁺/Mn²⁺): As A, but with 7 mM MgCl₂ and 0.15 mM MnCl₂.
- Reaction C (dNTP Imbalance): As A, but with 1 mM dATP/dTTP and 0.2 mM dCTP/dGTP.
- Reaction D (Combined): As B, but with the dNTP imbalance from C.
- Run PCR: 95°C for 2 min; [95°C for 30s, 55°C for 30s, 72°C for 1 min/kb] x 30 cycles; 72°C for 5 min.
- Clone and sequence 5-10 colonies from each reaction to calculate mutation frequency/kb.
- Set up a matrix of 4 reactions (50 µL each):
- A: Low diversity often stems from an insufficient mutation rate. To increase it, systematically adjust your epPCR conditions. Critical variables include:
Q2: How do I quantify and control the mutation rate in my epPCR?
- A: The mutation rate must be calibrated to your experimental goal. A low rate (1-2 mutations/kb) is for fine-tuning; a high rate (5-15 mutations/kb) is for broad exploration.
- Quantification: You must sequence a representative subset (at least 10-20 clones) from your initial library to determine the average number of mutations per kilobase.
- Control: Use the parameters in Table 1 as a starting guide. The mutation frequency is highly dependent on template sequence.
- A: The mutation rate must be calibrated to your experimental goal. A low rate (1-2 mutations/kb) is for fine-tuning; a high rate (5-15 mutations/kb) is for broad exploration.
Table 1: Effect of epPCR Parameters on Mutation Rate
| Parameter | Standard PCR Value | Low Mutation Rate (1-3/kb) | High Mutation Rate (8-15/kb) | Function |
|---|---|---|---|---|
| MgCl₂ | 1.5 mM | 1.5 - 3 mM | 5 - 7 mM | Stabilizes non-complementary base pairing. |
| MnCl₂ | 0 mM | 0 - 0.05 mM | 0.1 - 0.5 mM | Directly promotes misincorporation. |
| dNTPs | 0.2 mM each | 0.2 mM each | Imbalanced (e.g., 1 mM A/T, 0.2 mM C/G) | Imbalance reduces fidelity. |
| Taq Polymerase | 1.25 U/50µL | 1.25 U/50µL | 2.5 U/50µL | Higher enzyme concentration can increase error rate. |
| Template | 50-100 ng | 100 ng | <50 ng | Limits WT amplification. |
FAQ Category 2: Cloning & Assembly Efficiency
Q3: I am getting very low colony counts after Gibson Assembly/NEBuilder cloning of my PCR library. What should I check?
- A: Low efficiency in seamless cloning is typically due to suboptimal insert or vector preparation.
- Insert Purity: Purify the epPCR product twice (e.g., PCR cleanup kit followed by gel extraction for correct size). Residual primers and nucleotides inhibit assembly.
- Insert:Vector Ratio: Perform a molar ratio gradient. Test ratios from 2:1 to 10:1 (insert:vector). The ideal ratio is often 5:1.
- Vector Digestion: Ensure your linearized vector is completely digested and phosphatase-treated (e.g., with CIP or SAP) to prevent re-circularization. Run a small amount on a gel to confirm a single, clean band.
- Overlap Length: For methods like Gibson Assembly, ensure overlaps are 15-40 bp and have a Tm > 48°C. Use a calculator to design primers with optimal overlaps.
- Protocol: Gibson Assembly Optimization
- Prepare gel-purified insert(s) and vector.
- Set up a 20 µL assembly reaction with a molar ratio gradient:
- Tube 1: 2:1 Insert:Vector
- Tube 2: 5:1 Insert:Vector
- Tube 3: 10:1 Insert:Vector
- Use 2x Gibson/NEBuilder Master Mix. Incubate at 50°C for 15-60 minutes.
- Transform 2-5 µL into high-efficiency competent cells (>1 x 10⁸ cfu/µg). Plate and compare colony counts.
- A: Low efficiency in seamless cloning is typically due to suboptimal insert or vector preparation.
Q4: My final library has a high percentage of wild-type or empty vector clones. How can I reduce this background?
- A: This indicates inefficient cloning or selection.
- Use Positive Selection: Employ vectors with antibiotic resistance genes that are only restored upon successful insertion (e.g., α-complementation for blue/white screening or restriction-site elimination).
- DpnI Digestion: Treat your epPCR product with DpnI (post-PCR, pre-purification) to digest the methylated template plasmid, dramatically reducing parental background.
- Optimize Antibiotic Concentration: For libraries, use a higher than normal antibiotic concentration (e.g., 1.5x standard) to slow the growth of non-recombinants or cells with poorly expressed inserts.
- A: This indicates inefficient cloning or selection.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Library Construction |
|---|---|
| Taq DNA Polymerase | Standard polymerase for epPCR due to its inherent lack of 3'→5' exonuclease proofreading activity, allowing misincorporation of nucleotides. |
| MnCl₂ Solution | Critical additive for epPCR. Manganese ions destabilize polymerase fidelity, increasing the rate of base misincorporation. |
| DpnI Restriction Enzyme | Cuts methylated DNA. Used post-epPCR to selectively digest the original, bacterially-produced plasmid template, reducing wild-type background. |
| Gel Extraction Kit | For precise size-selection and purification of PCR inserts and linearized vectors, removing primers, enzymes, and mis-sized products. |
| Gibson Assembly / NEBuilder Master Mix | Contains exonuclease, polymerase, and ligase in an optimized buffer for seamless, one-pot assembly of multiple DNA fragments with homologous overlaps. |
| High-Efficiency Electrocompetent Cells | Essential for achieving large library sizes (>10⁶ clones). Electroporation offers significantly higher transformation efficiency than chemical methods. |
| Phosphatase (CIP/SAP) | Removes 5' phosphate groups from linearized vectors to prevent self-ligation, a major source of empty vector background. |
Visualizations
Diagram 1: Workflow for Bias-Minimized Mutagenic Library Construction
Diagram 2: Parameters Influencing epPCR Mutational Bias
Solving the Skew: Diagnostic and Corrective Strategies for Problematic Libraries
Troubleshooting Guides & FAQs
Q1: My post-error-prone PCR library shows significantly lower sequence diversity than expected based on the theoretical mutation rate. How do I start diagnosing this? A: Begin by mapping the potential failure point. First, run the purified PCR product on a high-sensitivity bioanalyzer or gel. If the smear is of the expected size, the initial PCR may be fine. Next, quantify the PCR product before and after your cloning step (e.g., ligation/transformation or Gibson assembly/E. coli transformation). A drop of >100-fold in recoverable clones vs. input DNA suggests a cloning bottleneck. If clone numbers are high, proceed to Sanger sequence 20-30 individual clones from the post-transformation pool before screening. Low diversity here points to PCR bias or cloning bias. High diversity here but low diversity after your functional screen indicates a screening failure.
Q2: What specific signs suggest the problem is PCR bias? A: Key indicators of PCR bias include: 1) Skewed nucleotide changes: A strong bias towards specific transitions (e.g., A•T → G•C) over transversions in your final sequence data. 2) Position-specific trends: Mutations are overwhelmingly clustered in certain regions of the amplicon, while others are "cold spots." 3) Homogeneous product size: The PCR product appears as a tight, single band instead of a heterogeneous smear (for high-mutation-rate epPCR). 4) Replicate consistency: The same few dominant sequences appear across independent PCR reactions.
Q3: I suspect a cloning bottleneck. What are the common culprits in library construction? A: The primary culprits are:
| Culprit | Effect | Diagnostic Test |
|---|---|---|
| Inefficient Ligation/Gibson Assembly | Low yield of full-length insert-vector constructs. | Run assembly product on gel; look for shift to higher molecular weight. Use positive control DNA. |
| Overly Restrictive Vector | Vector backbone may contain toxic elements or promoters. | Transform the empty vector backbone alone; assess colony count. Use a dedicated library construction vector. |
| Inefficient Competent Cells | Low transformation efficiency drastically under-samples the library. | Perform a control transformation with a known amount of supercoiled plasmid (e.g., 10 pg pUC19). Efficiency should be >1x10^7 cfu/µg for library work. |
| Toxic Inserts/Expression Leakage | Clones expressing toxic mutant proteins are lost during growth. | Clone a non-mutated version of your gene. If this also yields low counts, toxicity is likely. Use tight repression (e.g., T7 lacO, arabinose promoter) until screening. |
Q4: My cloning yield seems good, but my functional screen (e.g., phage display, FACS, colony assay) yields only a handful of unique variants. Is this a screening failure? A: Possibly. First, quantify the screening bottleneck. Calculate the ratio of screened clones to library diversity. If you screened 10^5 cells but your pre-screening library had 10^8 unique members, you've sampled only 0.1% of diversity—this is a sampling bottleneck, not a failure. A true screening failure occurs when the assay conditions are too stringent or misconfigured, eliminating all but a few variants. Test your screen with a known mix of positive and negative control clones to verify it can recover the positives at the expected frequency.
Q5: How can I experimentally distinguish between these three failure modes? A: Follow this sequential protocol:
Protocol: Diagnostic Workflow for Low Diversity
- Input Material Analysis: Gel purify the error-prone PCR product. Perform High-Throughput Sequencing (Illumina MiSeq) on the PCR product directly. This data reveals true mutation spectrum and diversity before any cloning.
- Post-Cloning, Pre-Screening Analysis: Pick 96 random colonies from the transformation plate (before any screening). Prepare a pooled plasmid mini-prep and submit for Illumina sequencing. Alternatively, sequence 20-30 clones individually by Sanger.
- Compare Datasets: Create a table comparing key metrics.
| Diversity Metric | Direct PCR-seq Data | Post-Cloning Pool-seq Data | Conclusion |
|---|---|---|---|
| Effective Unique Sequences | High (e.g., 10^5) | Low (e.g., 10^2) | Cloning Bottleneck |
| Mutation Spectrum (A•T→G•C %) | Balanced (~30%) | Skewed (>70%) | PCR Bias present |
| Mutation Distribution | Even across gene | Clustered in regions | PCR Bias present |
| Effective Unique Sequences | High | High | Problem is downstream (Screening Failure/Bottleneck) |
Q6: How can I mitigate PCR bias in error-prone PCR? A: Use the following experimental strategies:
- Polymerase Choice: Test different mutagenic polymerases (e.g., Mutazyme II, Taq Pol) as each has distinct bias profiles.
- Balanced Nucleotide Buffers: Use commercial kits (e.g., GeneMorph II) that adjust dNTP ratios to promote more balanced mutation rates.
- Template Reduction: Use the minimum amount of template DNA (e.g., 10^3 - 10^4 copies) to prevent wild-type carryover.
- Limited Cycling: Reduce PCR cycles to the minimum required for yield to avoid over-amplifying early biases.
- Fragmentation & Reassembly: Use methods like SHIPREC which fragment and reassemble mutated genes, breaking linkage biases.
Q7: What are key reagents for building a robust, diverse library? A:
Research Reagent Solutions Toolkit
| Item | Function | Example/Note |
|---|---|---|
| Balanced Error-Prone PCR Kit | Provides optimized buffer/dNTP mixes for more random mutagenesis. | GeneMorph II Random Mutagenesis Kit (Agilent) |
| High-Efficiency Cloning Vector | Vector with tight transcriptional control, lacking toxic elements, designed for library construction. | pET-29b(+) for T7 control; pBAD for arabinose control |
| Ultra-High Efficiency Competent Cells | E. coli strains with >1x10^8 cfu/µg transformation efficiency for maximum library representation. | NEB 10-beta, MegaX DH10B T1R, commercial electrocompetent cells |
| Next-Generation Sequencing Service | For deep diversity analysis of PCR products and cloned libraries. | Illumina MiSeq (2x300 bp) for full gene coverage. |
| Gel Extraction Kit (High Sensitivity) | For clean size-selection of heterogenous error-prone PCR products. | Qiagen QIAquick Gel Extraction, Zymoclean Gel DNA Recovery |
| Phusion High-Fidelity DNA Polymerase | For subsequent amplification of libraries with minimal added bias. | Used for "repair" or booster PCR after initial mutagenesis. |
Diagnostic Workflow Diagram
Diagnosing Low Diversity: A Troubleshooting Workflow
PCR Bias Mechanism Diagram
Mechanisms Leading to PCR Bias in Mutagenesis
Welcome to the Technical Support Center for addressing AT/GC bias in error-prone PCR mutagenesis. This guide provides troubleshooting and methodologies framed within the broader thesis of correcting mutagenesis bias to achieve uniform library diversity.
Troubleshooting Guides & FAQs
Q1: My error-prone PCR library shows a severe under-representation of AT-rich sequences. What is the primary cause and initial step? A1: This is a classic symptom of standard Taq DNA polymerase bias, which has a higher processivity for GC-rich templates. The initial step is to quantify your starting genomic template's AT/GC content. Use the following protocol:
- Protocol: Genomic Content Quantification
- Reagent: Purified genomic DNA template.
- Method: Perform a standard 50 µL PCR with primers flanking your target region using a high-fidelity polymerase (e.g., Q5). Purify the amplicon.
- Analysis: Use spectrophotometry (NanoDrop) to measure absorbance at 260nm. Calculate the ratio: %GC = (G + C) / (A + T + G + C) * 100. Alternatively, use bioinformatics tools (e.g., SeqKit stats) on Sanger sequencing data of the amplicon.
- Threshold: A template with >70% GC or <30% GC is considered "extreme" and requires rebalancing strategies.
Q2: How do I adjust nucleotide concentrations to rebalance bias, and what are the risks? A2: Manipulating dNTP ratios is a common strategy but requires precise optimization to avoid excessive error rate or PCR failure.
- For AT-rich genomes (to boost GC mutations): Increase the concentration of dCTP and dGTP relative to dATP and dTTP.
- For GC-rich genomes (to boost AT mutations): Increase the concentration of dATP and dTTP relative to dCTP and dGTP.
Protocol: dNTP Ratio Titration Experiment
- Prepare a master mix containing buffer, MgCl₂, primers, polymerase, and a fixed total dNTP concentration (e.g., 200 µM).
- Aliquot the master mix and spike in different ratios of biased dNTPs. A standard unbalanced condition might use a 1:1:8:8 ratio (dATP:dTTP:dCTP:dGTP for AT-rich targets, or the inverse for GC-rich).
- Run the error-prone PCR program.
- Clone and sequence 50-100 colonies per condition to analyze mutation spectrum and frequency.
Risk: Extreme imbalances can lead to misincorporation-driven polymerase stalling and reduced library size.
Q3: Which polymerase blends are best suited for extreme genomic content, and how do I choose? A3: No single polymerase is perfect. Blends are often used. See the comparison table below.
Q4: My optimized protocol still yields skewed libraries. What advanced strategy can I implement? A4: Consider tri-nucleotide phosphorothioate analogs (e.g., dNTPαS). These are incorporated but then poorly extended, forcing the polymerase to incorporate the next nucleotide from a biased pool, subtly shifting the mutation spectrum. This is a more advanced and costly reagent strategy.
Data Presentation
Table 1: Polymerase & Reagent Solutions for AT/GC Bias Correction
| Reagent/Enzyme | Primary Function | Best For | Key Consideration |
|---|---|---|---|
| Mutazyme II | Engineered blend for uniform mutation distribution. | General-purpose bias reduction. | Proprietary blend; optimal with 0.1-0.5 mM Mn²⁺. |
| Taq + Pol I (Klenow) | Taq creates lesions, Klenow extends with lower bias. | AT-rich templates. | Requires two-step addition; ratio optimization needed. |
| GeneMorph II | Mutazyme-based, optimized for random mutagenesis. | GC-rich templates. | Works across a wide GC range (30-80%). |
| Biased dNTP Pools | Unbalanced nucleotide concentrations. | Fine-tuning specific mutation types. | Requires careful titration to avoid PCR failure. |
| Mn²⁺ (Manganese) | Replaces Mg²⁺ to reduce polymerase fidelity. | Increasing overall error rate. | Can exacerbate sequence bias if used alone. |
| Tri-nucleotide dNTPαS | Causes transient chain termination after biased incorporation. | Advanced spectral shifting. | Expensive; requires specialized synthesis. |
Table 2: Quantitative Outcomes of Different Rebalancing Strategies
| Strategy | Target GC% | Mutation Rate (mut/kb) | AT→GC:GC→AT Mutation Ratio | Library Coverage (%)* |
|---|---|---|---|---|
| Standard Taq/Mn²⁺ | 25% (AT-rich) | 12.5 | 0.4 : 1 | ~45% |
| Biased dNTP (8:8:1:1) | 25% (AT-rich) | 8.2 | 0.9 : 1 | ~78% |
| Mutazyme II Blend | 25% (AT-rich) | 10.1 | 1.1 : 1 | ~92% |
| Standard Taq/Mn²⁺ | 75% (GC-rich) | 9.8 | 3.2 : 1 | ~35% |
| Biased dNTP (1:1:8:8) | 75% (GC-rich) | 7.5 | 1.5 : 1 | ~82% |
| GeneMorph II Blend | 75% (GC-rich) | 11.3 | 1.3 : 1 | ~88% |
*Theoretical coverage of sequence space in the target region.
Experimental Protocols
Core Protocol: Error-Prone PCR with Bias Correction
Objective: Generate a mutagenic library with uniform distribution across an extreme-GC or extreme-AT template. Materials: See "The Scientist's Toolkit" below. Procedure:
- Template Quantification: Determine the precise AT/GC% of your target amplicon (see Protocol Q1).
- Strategy Selection: Based on Table 1 & 2, select a polymerase/blend and decide if dNTP bias is needed.
- Primary PCR Setup:
- 50 µL Reaction:
- 10-50 ng genomic or plasmid template.
- 1X proprietary buffer (supplied with enzyme).
- dNTPs: Either standard 0.2 mM each or a biased mix (e.g., 0.05 mM dATP/dTTP, 0.35 mM dCTP/dGTP for GC-rich targets).
- Divalent Cations: 0.3-0.5 mM Mn²⁺ (critical for error generation). Adjust Mg²⁺ as per enzyme guidelines.
- 0.3 µM each forward and reverse primer.
- 1-2 units of selected mutagenic polymerase blend.
- Thermocycling: Standard 3-step PCR (94°C denaturation, 50-55°C annealing, 72°C extension) for 25-30 cycles.
- 50 µL Reaction:
- Purification: Purify the PCR product using a spin column or magnetic beads.
- Analysis: Clone into your desired vector, transform, and sequence a statistically significant number of clones (≥50) to calculate mutation rate and spectrum using tools like Enrich.
Mandatory Visualization
Title: Workflow for Rebalancing Error-Prone PCR Bias
Title: Key Strategies for Correcting PCR Mutagenesis Bias
The Scientist's Toolkit
Research Reagent Solutions
| Item | Function in Bias Correction |
|---|---|
| Mutazyme II DNA Polymerase | Proprietary enzyme blend designed to produce a uniform mutation spectrum independent of sequence context. |
| GeneMorph II Random Mutagenesis Kit | Optimized system (enzyme, buffer, control template) for mutating difficult, GC-rich targets. |
| Unbalanced dNTP Set | Separate dATP, dTTP, dCTP, dGTP stocks for creating custom biased nucleotide ratios. |
| Manganese Chloride (MnCl₂) | The critical divalent cation for reducing polymerase fidelity and enabling error-prone PCR. |
| High-Fidelity Polymerase (e.g., Q5) | Used for initial template amplification and library amplification post-mutagenesis without introducing extra errors. |
| Next-Generation Sequencing (NGS) Service | Essential for deep analysis of final library mutation spectrum and uniformity. |
| Tri-nucleotide Phosphorothioate Analogs | Specialized nucleotides for advanced, forced misincorporation strategies. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: Our error-prone PCR library shows an unexpectedly high frequency of frameshift mutations and non-functional clones. What could be the cause and how can we mitigate this? A: This is a classic symptom of excessive mutational load. The primary cause is often an overly high mutation rate, typically from using too much Mn2+ or unbalanced dNTP concentrations in the error-prone PCR protocol.
- Troubleshooting Steps:
- Quantify Mutation Frequency: Sequence 20-50 random clones to determine the average number of mutations per gene. Aim for 1-4 amino acid substitutions per gene, depending on length.
- Adjust Protocol: Reduce MnCl2 concentration incrementally (try 0.05-0.2 mM steps). Ensure standard Mg2+ is present (1-2 mM).
- Use dNTP Analogues: Replace dCTP and dTTP with 8-oxo-dGTP and dPTP at a ratio of 10:1 (standard:analogue) to promote transition mutations over transversions, which are less likely to be catastrophic.
- Post-PCR Size Selection: Use agarose gel electrophoresis to isolate the correctly sized product, filtering out larger indels.
Q2: How can we design a screening strategy to efficiently identify functional variants amidst a high background of deleterious mutations? A: Implement a tiered screening funnel to manage load.
- Workflow:
- Primary Screen (High-Throughput): Use a functional complementation or survival assay (e.g., antibiotic resistance, auxotrophy) to quickly eliminate null mutants. This removes frameshifts and severe deleterious mutations.
- Secondary Screen (Medium-Throughput): Apply a quantitative assay (e.g., fluorescence, enzymatic activity in colonies) to rank functional variants.
- Tertiary Validation (Low-Throughput): Sequence top performers and characterize purified proteins.
Q3: What bioinformatic tools can help predict and filter deleterious mutations in silico before experimental screening? A: Several tools can prioritize variants.
- SIFT & PROVEAN: Predict whether an amino acid substitution affects protein function.
- FoldX: Predicts the effect of mutations on protein stability (ΔΔG).
- Action: After sequencing your initial library, filter out variants predicted as deleterious or highly destabilizing (e.g., FoldX ΔΔG > 2-3 kcal/mol) to focus resources on promising candidates.
Q4: The mutational spectrum from our error-prone PCR is biased. How can we achieve a more uniform distribution of mutations? A: Standard error-prone PCR has known sequence context biases.
- Solution:
- Use Commercial Kits: Kits like the GeneMorph II Random Mutagenesis Kit (Agilent) or the Diversify PCR Mutagenesis Kit (Takara) employ engineered mutase enzymes for more random incorporation.
- Combine Methods: Use DNA shuffling or staggered extension process (StEP) after error-prone PCR to recombine beneficial mutations and separate them from deleterious ones.
Table 1: Effect of Mn2+ Concentration on Mutation Rate & Quality
| [MnCl2] (mM) | Avg. Mutations/kb | % Functional Clones | % Frameshift Indels |
|---|---|---|---|
| 0.05 | 2-4 | 65% | <2% |
| 0.10 | 4-8 | 40% | 5% |
| 0.20 | 8-16 | 15% | 15% |
| 0.50 | 16+ | <5% | >25% |
Table 2: Comparison of Mutagenesis Methods
| Method | Avg. Mutation Rate | Bias | Best for |
|---|---|---|---|
| Standard epPCR (Mn2+) | Medium-High | High (AT bias) | Low-diversity exploration |
| epPCR with dNTP Analogues | Medium | Reduced | More transitions, fewer stop codons |
| Commercial Random Mutagenesis Kits | Tunable (Low-High) | Low | Uniform random libraries |
| Site-Saturation Mutagenesis | Targeted (1 codon) | None | Deep analysis of specific sites |
Experimental Protocols
Protocol 1: Optimized Error-Prone PCR for Controlled Mutational Load Objective: Generate a library with 1-4 amino acid changes per 1kb gene. Reagents: Taq DNA Polymerase, 10X Buffer (no Mg2+), 25 mM MgCl2, 10 mM dNTPs (including analogues if desired), 10 mM MnCl2, template DNA, primers. Procedure:
- Prepare a 50 µL reaction on ice:
- 5 µL 10X Taq Buffer (Mg-free)
- Template DNA (50-100 ng)
- Primers (0.2 µM each)
- 0.2 mM each dATP, dGTP
- 0.2 mM each dCTP, dTTP (For biased spectrum, replace with 0.18 mM dCTP + 0.02 mM 8-oxo-dGTP and 0.18 mM dTTP + 0.02 mM dPTP)
- 2.0 mM MgCl2
- 0.1 mM MnCl2 (Critical: Titrate from 0.05-0.15 mM for optimization)
- 2.5 U Taq DNA Polymerase
- Nuclease-free water to 50 µL.
- PCR Cycling:
- 95°C for 2 min.
- 25 Cycles: 95°C for 30 sec, 55-60°C (primer-specific) for 30 sec, 72°C for 1 min/kb.
- 72°C for 5 min.
- Purify PCR product using a spin column.
- Size Selection: Run product on an agarose gel. Excise the band corresponding to the correct size to remove larger/smaller indel products.
- Gel-purify the size-selected DNA. This is your mutagenic library for cloning.
Protocol 2: Primary Functional Screen via Complementation Objective: Rapidly eliminate non-functional clones. Procedure:
- Clone the error-prone PCR library into an appropriate expression vector.
- Transform the ligation into a complementation host strain (e.g., for an enzyme in a metabolic pathway, use an auxotrophic strain requiring that pathway's product).
- Plate transformants onto selective media lacking the essential metabolite.
- Only clones expressing functional variants will grow. Pool these colonies for plasmid extraction and secondary screening.
Mandatory Visualizations
Title: Functional Screening Funnel for Mutant Libraries
Title: Optimized epPCR & Size Selection Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Managing Mutational Load
| Reagent / Kit | Function & Rationale |
|---|---|
| Taq DNA Polymerase | Lacks 3'→5' exonuclease proofreading, allowing nucleotide misincorporation during error-prone PCR. |
| Manganese Chloride (MnCl2) | Critical divalent cation. Causes polymerase misincorporation by increasing error rate. Must be titrated precisely. |
| 8-oxo-dGTP & dPTP | Nucleotide analogues. Bias mutations toward less deleterious transition mutations (AG, CT) over transversions. |
| GeneMorph II Random Mutagenesis Kit (Agilent) | Uses engineered mutase enzymes for tunable, more uniform mutation distribution with low frameshift incidence. |
| Diversify PCR Mutagenesis Kit (Takara) | Provides a blend of nucleotides and polymerase for controlled, random mutagenesis without extreme bias. |
| FoldX Protein Engineering Software | In silico tool to predict stability change (ΔΔG) of mutations. Filters destabilizing variants pre-screening. |
| Size-Selective Agarose Gels | Critical physical method to remove PCR products with large insertions/deletions (indels) that cause frameshifts. |
| Complementation Host Strains | Genetically engineered cells (E. coli, yeast) that require target protein function for survival on selective media. Enables high-throughput primary functional screen. |
Troubleshooting Poor Amplification Yield and Template Re-Amplification Issues
Troubleshooting Guides & FAQs
FAQ 1: Why is my amplification yield low or non-existent? This is often due to suboptimal reaction conditions, inhibitor presence, or poor template quality. A systematic approach is required to isolate the variable.
FAQ 2: What are the primary risks of re-amplifying a PCR product? Re-amplification significantly increases mutagenesis bias in error-prone PCR by selectively amplifying sequences that were favored in the first round, compounding errors and skewing the mutant library diversity. It should be avoided when studying unbiased mutagenesis.
FAQ 3: How can I troubleshoot a failed PCR before considering re-amplification? Follow a stepwise checklist: verify template integrity and concentration, check primer design and annealing temperatures, ensure reagent activity (especially polymerase), and optimize Mg²⁺ concentration and cycling parameters.
FAQ 4: What are the best practices to avoid the need for re-amplification? Optimize the primary reaction using a gradient PCR, include positive and negative controls, use high-fidelity or dedicated error-prone polymerases with appropriate buffers, and accurately quantify template DNA.
Detailed Troubleshooting Guide: Low Yield
- Problem: No band or faint band observed on gel post-PCR.
Solution Path:
- Control Check: Run a positive control (known template/primer set) with your master mix. If it fails, your reagents (polymerase, dNTPs, buffer) are likely degraded.
- Template Check:
- Verify concentration and purity (A260/A280 ratio of ~1.8).
- Run template on a gel to check for degradation.
- Dilute template to reduce potential inhibitors.
- Primer Check:
- Verify primer concentration (typical final conc. 0.1–1 µM).
- Check for primer-dimer formation in a no-template control.
- Re-calculate primer Tm and use a gradient PCR to find optimal annealing.
- Reaction Condition Check:
- Increase MgCl₂ concentration in 0.5 mM increments (Mg²⁺ is a cofactor for polymerase).
- Increase the number of cycles cautiously (for error-prone PCR, typically 25-30 cycles; exceeding this increases bias).
- Ensure the denaturation temperature and time are sufficient for your template.
Experimental Protocol: Optimizing Mg²⁺ Concentration for Error-Prone PCR
- Prepare a standard error-prone PCR master mix, omitting MgCl₂.
- Aliquot the master mix into 6 tubes.
- Add MgCl₂ from a stock solution to achieve final concentrations of 1.0, 1.5, 2.0, 2.5, 3.0, and 3.5 mM.
- Add template and primers to each tube.
- Run the PCR using your standard cycling protocol.
- Analyze products by agarose gel electrophoresis to determine the concentration yielding the strongest, correct product band with minimal non-specific amplification.
Quantitative Data Summary
Table 1: Common PCR Issues and Their Quantitative Adjustments
| Issue | Suspect Variable | Typical Starting Range | Optimization Step |
|---|---|---|---|
| Low Yield | Template Amount | 0.1–100 ng (genomic) | Titrate in 10-fold dilutions |
| Low Yield | MgCl₂ Concentration | 1.5 mM (standard) | Titrate from 1.0 to 3.5 mM in 0.5 mM steps |
| Low Yield | Annealing Temperature | Primer Tm ± 5°C | Run a gradient 5°C below to 5°C above calculated Tm |
| Non-Specific Bands | Annealing Temperature | As above | Increase temperature incrementally within gradient |
| Primer-Dimer Formation | Primer Concentration | 0.1–1 µM | Reduce concentration to 0.1–0.2 µM |
Table 2: Impact of Re-Amplification on Mutagenesis Bias
| Experimental Step | Average Mutation Frequency (per kb) | Library Diversity (% unique variants) | Risk of Over-representing Early Mutants |
|---|---|---|---|
| Primary EP-PCR (Cycle 25) | 5-15 | High (~80-95%) | Low |
| First Re-Amplification (Total ~50 cycles) | 15-40 | Moderate (~50-70%) | High |
| Second Re-Amplification (Total ~75 cycles) | 40+ | Low (<30%) | Very High |
Workflow Diagram
Title: Troubleshooting Workflow for Poor PCR Yield
Error-Prone PCR Bias Pathway
Title: How Re-Amplification Compounds PCR Bias
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Error-Prone PCR & Troubleshooting
| Reagent / Material | Function in Experiment | Key Consideration for Yield/Bias |
|---|---|---|
| Error-Prone Polymerase Blend (e.g., Mutazyme II, Taq Pol with Mn²⁺) | Introduces controlled random mutations during amplification. | Specific buffer and Mg²⁺/Mn²⁺ ratios are critical for yield and mutation rate. |
| MgCl₂ & MnCl₂ Stock Solutions | Divalent cation cofactors. Mn²⁺ increases error rate. | Concentration must be precisely optimized; slight changes drastically affect yield and fidelity. |
| High-Purity dNTP Mix | Building blocks for DNA synthesis. | Imbalanced dNTP pools can increase error rate but also cause premature termination. |
| PCR Grade Water | Solvent for master mixes. | Must be nuclease-free to prevent degradation of primers/template. |
| Thermostable Polymerase (Standard Taq) | For diagnostic/setup PCRs during troubleshooting. | Use to test reaction components without the variable of mutagenesis. |
| Gel Extraction & Clean-up Kit | Purifies PCR product from agarose gel or reaction mix. | Essential for removing primers, enzymes, and salts before any downstream application. Do not use to enable re-amplification. |
| Quantitation Spectrophotometer (e.g., Nanodrop) | Accurately measures DNA template concentration and purity. | Prevents under- or over-loading template, a common cause of failure. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My error-prone PCR (epPCR) library shows drastically lower diversity than expected based on theoretical calculations. What could be the cause?
A: This is a common manifestation of mutagenic bias. The calculated diversity assumes equal probability for all nucleotide substitutions, which is rarely true with standard epPCR protocols.
- Primary Causes & Solutions:
- Uneven Nucleotide Incorporation: Standard polymerases (e.g., Taq) often have a bias against incorporating dNTP analogs like 8-oxo-dGTP or dPTP.
- Solution: Use a polymerase blend optimized for epPCR (e.g., Mutazyme II, GeneMorph II). These are engineered for more balanced mutation rates.
- Skewed Mutation Spectrum: The type of dNTP analog and its concentration relative to natural dNTPs heavily influences the transition/transversion ratio.
- Solution: Titrate the Mn²⁺ concentration and the ratio of mutagenic to natural dNTPs. Refer to the table below for standardized adjustments.
- Template GC-content: High GC-content templates can lead to "cold spots" with minimal mutation.
- Solution: Incorporate a small amount of dITP (deoxyinosine triphosphate) to destabilize GC pairs, or use PCR additives like DMSO or betaine.
- Uneven Nucleotide Incorporation: Standard polymerases (e.g., Taq) often have a bias against incorporating dNTP analogs like 8-oxo-dGTP or dPTP.
Q2: After screening my mutant library, I find an overwhelming number of variants that are insoluble or misfolded, even with low target mutation rates. How can I better preserve protein folding?
A: This indicates a failure to constrain the explored sequence space to regions compatible with the protein's structural framework.
- Primary Causes & Solutions:
- Unconstrained Randomness: Fully random mutagenesis, especially at buried residues, disrupts hydrophobic cores and key stabilizing interactions.
- Solution: Implement site-saturation mutagenesis (SSM) focused on surface loops or active site peripheries instead of whole-gene epPCR. Use structure-guided libraries.
- Ignoring Covariance: Natural evolution preserves pairs of residues that co-vary (e.g., salt bridges). Random single-point mutations break these.
- Solution: Use coupled mutation libraries based on statistical coupling analysis (SCA) or direct evolutionary coupling analysis (ECA) of homologous sequences.
- Lack of Functional Pre-screening: Expression in a non-functional context (e.g., inclusion bodies) wastes screening effort.
- Solution: Employ phage or yeast display coupled with a binding step (even to the wild-type antigen) to pre-filter for properly folded variants before detailed activity assays.
- Unconstrained Randomness: Fully random mutagenesis, especially at buried residues, disrupts hydrophobic cores and key stabilizing interactions.
Q3: How can I accurately measure and report the actual mutational bias in my epPCR library for my thesis methodology section?
A: You must sequence a random subset of the library before selection to establish the baseline mutation frequency and spectrum.
- Experimental Protocol: Deep Sequencing Analysis of Library Bias
- Library Preparation: Perform your epPCR reaction. Purify the product.
- Sample for NGS: Clone the library into a simple vector and transform E. coli. Pick at least 50-100 colonies for Sanger sequencing, or prepare amplicons for high-throughput NGS (e.g., Illumina MiSeq).
- Data Analysis:
- Align sequences to the parent gene.
- Calculate: Average Mutation Frequency (total mutations / total bp sequenced).
- Categorize mutations: Transitions (AG, CT) vs. Transversions (all others).
- Create a position-specific map of mutations to identify "hot" and "cold" spots.
- Reporting: The mutation distribution table (see below) is essential for your thesis to document the bias.
Table 1: Comparison of Common Error-Prone PCR Methods and Their Biases
| Method / Reagent Kit | Typical Mutation Rate (mutations/kb) | Key Bias | Optimal for... | Folding Preservation Strategy |
|---|---|---|---|---|
| Standard Taq + Mn²⁺ | 1-10 | High AT→GC bias; hotspot prone | Low-complexity exploration | Poor; often requires subsequent solubility screening. |
| Mutazyme I/II (Agilent) | 1-16 | More balanced spectrum than Taq | General purpose, medium-length genes | Moderate; more even distribution can help. |
| GeneMorph II (Agilent) | 0.1-16 (adjustable) | Low bias, random | Tunable libraries for directed evolution | Good when tuned for low-to-medium rates. |
| TriNucleotide Mutagenesis (TRIM) | Site-specific | Virtually no codon bias | Saturation mutagenesis at defined sites | Excellent; allows exclusion of stop codons and non-sense mutations. |
Table 2: Analysis of a Hypothetical epPCR Library (NGS Data)
| Metric | Value | Comment |
|---|---|---|
| Average Mutation Frequency | 4.7 mutations/kb | Within target range (4-5/kb). |
| Transition : Transversion Ratio | 3.2 : 1 | High bias towards transitions (expected for Mn²⁺/Taq). |
| Most Frequent Substitution | A → G (28%) | Confirms strong polymerase bias. |
| % Sequences with Stop Codons | 12% | Explains loss of functional clones. |
| Conserved Position Mutation Rate | <0.1% | Indicates successful structural constraint via primer design. |
Experimental Protocols
Protocol: Structure-Guided Saturation Mutagenesis to Preserve Folding
Objective: To create a focused mutant library by randomizing only solvent-exposed, flexible residues while keeping structurally critical residues fixed.
Materials: See "The Scientist's Toolkit" below. Workflow:
- Identify Target Positions: Using your protein's crystal structure or a reliable homology model (e.g., from AlphaFold2), select residues with >40% solvent accessibility located in loops or flexible regions.
- Design Primers: For each target position, design degenerate primers using the NNK degeneracy (N = A/T/G/C; K = G/T). This encodes all 20 amino acids and one stop codon.
- PCR:
- Set up a QuikChange-style PCR reaction for each site:
- Template DNA: 50 ng plasmid.
- Primers: 125 ng each forward and reverse degenerate primer.
- Use a high-fidelity polymerase (e.g., PfuUltra II) to avoid secondary mutations.
- Thermocycling: 95°C 2 min; [95°C 30s, 55°C 1 min, 68°C X min (1 min/kb plasmid length)] x 18 cycles; 68°C 10 min.
- Set up a QuikChange-style PCR reaction for each site:
- DpnI Digestion: Treat PCR product with DpnI (37°C, 1 hr) to digest methylated parent template.
- Transform & Pool: Transform into competent E. coli, plate, and pool at least 5x library size colonies for plasmid extraction to create your final, folded-enriched library.
Mandatory Visualization
Diagram 1: Workflow for Bias-Aware Directed Evolution
Diagram 2: Key Factors in Functional Protein Space Optimization
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Bias-Optimized Mutagenesis
| Item (Vendor Examples) | Function in Experiment | Key Consideration |
|---|---|---|
| GeneMorph II Random Mutagenesis Kit (Agilent) | Provides optimized polymerase and buffer system for tunable, low-bias epPCR. | Allows control over mutation frequency via input DNA amount. |
| Phusion or Q5 High-Fidelity DNA Polymerase (NEB) | For site-directed/saturation mutagenesis without introducing secondary errors. | Critical for structure-guided library construction. |
| NNK Degenerate Codon Primers | Encodes all 20 amino acids + 1 stop codon in a single saturation mutagenesis primer. | Standard for focused library design. |
| Crystal Structure or AlphaFold2 Model | Identifies solvent-exposed, mutable residues vs. buried, conserved ones. | Foundation for structure-guided design. |
| Yeast Surface Display Kit (e.g., pYD1) | Enables folding pre-selection by displaying properly folded proteins on yeast surface. | Filters out misfolded variants before functional screening. |
| MiSeq Reagent Kit v3 (600-cycle) (Illumina) | For deep sequencing of pre-selection libraries to empirically quantify bias. | Essential for accurate thesis methodology reporting. |
| DMSO or Betaine (Sigma-Aldrich) | PCR additives that help amplify high-GC templates uniformly, reducing "cold spots". | Improves library uniformity. |
Beyond the Protocol: Validating Library Quality and Benchmarking Against Alternative Techniques
Troubleshooting Guides and FAQs
Q1: Our NGS library prepared from error-prone PCR (epPCR) products shows extremely low diversity and high duplication rates. What could be the cause and how can we fix it? A: This is a common issue stemming from insufficient input diversity or over-amplification during the NGS library preparation PCR. To address this: 1) Ensure your initial epPCR reaction generates a complex library by titrating Mn2+ concentration and cycle number. 2) For NGS library prep, use a high-fidelity polymerase and minimize the number of amplification cycles (typically 4-8 cycles). 3) Quantify the library using a fluorometric method (e.g., Qubit) and pool multiple, independent epPCR reactions before library prep to increase complexity.
Q2: We observe a high background of wild-type sequence reads in our deep sequencing data, masking low-frequency variants. How can we enrich for mutant sequences? A: High wild-type background often indicates low mutagenesis efficiency. First, optimize your epPCR protocol (see Table 1). If optimization fails, you can use a physical or enzymatic enrichment step post-epPCR but prior to NGS library prep. A recommended method is Nuclease-based Degradation of Wild-type Templates: Digest the epPCR product with a restriction enzyme or DpnI (which cuts methylated, template DNA) to selectively degrade the original plasmid template, enriching for newly synthesized mutant strands.
Q3: The mutation spectrum from our deep sequencing data shows a strong bias towards specific transitions (e.g., A•T → G•C). Is this expected, and how do we account for it in our analysis? A: Yes, bias is intrinsic to epPCR. Taq polymerase, especially with Mn2+, has known sequence-context-dependent mutation preferences. This must be accounted for when interpreting mutational spectra for thesis research on mutagenesis bias. The solution is to incorporate bias correction factors in your bioinformatics pipeline. Generate a control dataset by sequencing a non-functional region subjected to the same epPCR conditions to define the baseline biochemical bias of your system, which can then be used to normalize your experimental variant data.
Q4: Bioinformatics analysis is overwhelming. What are the essential steps to go from raw NGS reads to a reliable mutation frequency and spectrum table? A: A robust, minimal pipeline is required for gold-standard validation. Below is a core protocol.
Experimental Protocol: Core Bioinformatics Workflow for epPCR NGS Data Analysis
- Demultiplexing & QC: Use
bcl2fastqorbcl-convert(Illumina) to generate FASTQ files. Assess quality withFastQC. - Read Trimming & Filtering: Use
Trimmomaticorfastpto remove adapters and low-quality bases (threshold: Q20). - Alignment: Map reads to the reference wild-type sequence using a sensitive aligner like
BWA-MEMorBowtie2. Generate SAM/BAM files. - Processing Alignment: Sort and index BAM files (
samtools sort,samtools index). Mark duplicates if necessary (samtools markdup). - Variant Calling: For epPCR, use a tool that models amplicon data and handles PCR errors, such as
LoFreqorVarScan2(with--min-var-freqset appropriately, e.g., 0.1%). - Filtering & Annotation: Filter variants by depth (e.g., minimum 1000x coverage), strand bias, and quality score. Annote functional impact with
SnpEff. - Spectrum & Distribution: Use custom scripts (Python/R) to compile mutation types (C→T, A→G, etc.) and their spatial distribution along the gene.
Q5: How much sequencing depth is truly required for reliable detection of variants in an epPCR library? A: The required depth depends on the complexity of your library and the lowest variant frequency you need to detect confidently. For gold-standard validation aiming to capture the full spectrum, follow the guidelines in Table 1.
Table 1: Key Quantitative Parameters for epPCR NGS Validation
| Parameter | Recommended Value | Rationale |
|---|---|---|
| Average Sequencing Depth | 1,000 - 5,000x per nucleotide | Ensures statistical power to detect variants at frequencies as low as 0.1-0.5%. |
| Minimum epPCR Library Size | >10^7 independent clones | Ensures sufficient theoretical diversity for deep sequencing sampling. |
| Mutagenesis Rate Target | 1-20 mutations/kb | Optimal range for generating single and double mutants without excessive deleterious load. |
| NGS Read Length | 2x the length of your amplicon | Allows for paired-end overlap, providing higher quality consensus for variant calling. |
| Variant Frequency Cut-off | 0.1% (with appropriate depth) | Balances sensitivity with false-positive rate from PCR/NGS errors. |
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for epPCR and NGS Validation
| Item | Function |
|---|---|
| Mutazyme II or Taq Polymerase with MnCl2 | The core enzyme for biased, low-fidelity synthesis during error-prone PCR. |
| High-Fidelity Polymerase (e.g., Q5, KAPA HiFi) | Essential for NGS library amplification to avoid introducing new errors during prep. |
| Magnetic Beads (SPRI) | For size selection and clean-up of epPCR and NGS library products. |
| Dual-Indexed NGS Library Prep Kit | Allows multiplexing of multiple samples. Illumina-compatible kits are standard. |
| dsDNA HS Assay Kit (Qubit) | Accurate quantification of DNA libraries for pooling and loading. |
| DpnI Restriction Enzyme | Selectively digests methylated parental DNA template post-epPCR to reduce wild-type background. |
| Validated Primers with Overhangs | Gene-specific primers with 5' overhangs compatible with your NGS library adapter system. |
Workflow and Pathway Visualizations
Title: epPCR to NGS Analysis Workflow for Mutagenesis Bias Thesis
Title: Bioinformatics Pipeline for Mutation Analysis
Title: Framework for Correcting PCR Mutagenesis Bias in Analysis
Troubleshooting Guides & FAQs
General Mutagenesis Issues
Q1: My error-prone PCR (epPCR) library shows extremely low diversity and a strong sequence bias. What could be the cause and how can I fix it?
- A: This is a common issue rooted in the inherent nucleotide bias of Taq polymerase and uneven mutational incorporation. To mitigate:
- Use a specialized epPCR kit: Replace standard Taq with a blend like Mutazyme II, which reduces sequence context bias.
- Optimize Mn²⁺ concentration: Titrate MnCl₂ (0.05-0.5 mM) alongside standard Mg²⁺. Too much Mn²⁺ increases error rate but also bias.
- Diversify nucleotide pools: Use biased dNTP mixes (e.g., lower [dATP, dTTP]) to encourage transversions.
- Consider orthogonal validation: Use a low-bias method like CRISPR-based mutagenesis (see Table 1) to target the specific region of interest as a control.
Q2: When using CRISPR-based mutagenesis for saturation editing, I'm getting low editing efficiency and high wild-type background. How do I improve this?
- A: This typically points to gRNA and repair template issues.
- Verify gRNA efficiency: Use a validated online tool (e.g., CRISPick) to score your gRNA for your organism. Ensure the PAM site is correctly identified.
- Optimize repair template (donor DNA) design: Use single-stranded oligodeoxynucleotides (ssODNs) with homology arms of 35-90 nt. Incorporate synonymous "blocking" mutations in the PAM sequence to prevent re-cutting.
- Check delivery ratios: For plasmid-based systems, maintain a 1:1:1 molar ratio of Cas9 plasmid, gRNA plasmid, and donor DNA. Increase the donor DNA amount if needed.
- Use a high-fidelity Cas9 variant: Reduces off-target effects that can compromise cell viability and library quality.
Q3: In Phage-Assisted Continuous Evolution (PACE), my phage titer drops to zero rapidly, halting evolution. What are the critical checkpoints?
- A: Phage loss is catastrophic. Troubleshoot systematically:
- Host cell viability: Ensure the E. coli host cells (e.g., S2060) are healthy and the accessory plasmid (AP) expressing the selection factor is functional.
- Lagoon dilution rate: Calculate the dilution rate precisely. If the lagoon dilutes faster than the phage replication rate under selection, phage will wash out. Start with a slower dilution (e.g., 1 lagoon volume per hour) and increase gradually.
- Selection stringency: The activity of the mutagenized gene on the AP must confer a clear replication advantage. Verify the selection circuit with a known functional protein control before starting evolution.
- Mutation rate: Ensure the mutagenesis plasmid (MP) is present and active. Low mutation rates prevent adaptive variants from arising in time.
Protocol-Specific Issues
Q4: What is a robust protocol for performing a controlled, low-bias epPCR reaction?
- A: Detailed epPCR Protocol (using a commercial kit for reliability)
- Reagent: GeneMorph II Random Mutagenesis Kit (Agilent).
- Setup: In a 50 µL reaction, combine:
- 10-100 ng of template DNA.
- 1x Mutazyme II reaction buffer.
- 0.5 µL Mutazyme II DNA polymerase.
- Forward and reverse primers (0.3 µM each).
- dNTP mix (provided).
- Nuclease-free water.
- Cycling Conditions:
- 95°C for 2 min (initial denaturation).
- 95°C for 30 sec (denaturation).
- Tm of primers for 30 sec (annealing). [Note: User must calculate based on primers].
- 72°C for 1 min/kb (extension).
- Repeat steps 2-4 for 25 cycles. [Note: Cycle number directly controls mutation rate].
- 72°C for 10 min (final extension).
- Post-PCR: Purify the product using a spin column before cloning.
Q5: What is a standard protocol for multiplexed CRISPR saturation mutagenesis in yeast?
- A: Detailed CRISPR Saturation Mutagenesis Protocol
- Step 1: Library Design. Design oligos to replace a target codon with an NNK degenerate codon (encodes all 20 amino acids) via Cas9 cutting. Clone oligo pool into a donor plasmid backbone.
- Step 2: Transformation. Co-transform competent yeast cells (e.g., S. cerevisiae) with:
- The donor plasmid library.
- A Cas9 expression plasmid.
- A plasmid expressing the target-specific gRNA.
- Step 3: Selection & Screening. Plate transformations on selective media. Screen colonies via diagnostic PCR and sequencing to confirm editing and library diversity.
- Step 4: Functional Assay. Subject the pooled library to your phenotypic selection (e.g., growth assay, fluorescence sorting).
Performance Benchmarking Data
Table 1: Comparative Benchmarking of Mutagenesis Methods
| Feature | Error-Prone PCR (epPCR) | CRISPR-based Saturation Mutagenesis | Phage-Assisted Continuous Evolution (PACE) |
|---|---|---|---|
| Theoretical Library Diversity | Very High (10^10-10^13) | Targeted - High per site (32-64) | Continuous, theoretically infinite |
| Practical Library Size | 10^6 - 10^8 variants | Limited by transformation efficiency (~10^7-10^9) | 10^10 - 10^12 phages per lagoon |
| Mutation Bias | High (Prone to AT>GC transitions) | Very Low (Controlled by oligo design) | Moderate (Dependent on MP) |
| Control Over Mutation Location | Random across gene | Precise to a single codon/region | Random across genome |
| Evolutionary Pressure | In vitro, no selection | In vivo, post-editing selection | Continuous, direct selection |
| Typical Timeframe (Cycle) | 3-5 days | 5-10 days | Weeks, but continuous |
| Key Advantage | Simple, large random libraries | Precision, low bias, combinatorial | Hands-off, explores vast fitness landscapes |
| Primary Limitation | Sequence bias, screening burden | Throughput per experiment limited | Complex setup, organism restriction |
Experimental Workflow Diagrams
Title: Standard epPCR Library Generation Workflow
Title: CRISPR-based Saturation Mutagenesis Protocol
Title: PACE Selection Logic and Continuous Evolution
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Mutagenesis Studies
| Reagent/Material | Function in Experiment | Example Product/Kit |
|---|---|---|
| Mutazyme II DNA Polymerase Blend | Reduces sequence bias in epPCR for more random mutagenesis. | Agilent GeneMorph II Random Mutagenesis Kit |
| NNK Degenerate Codon Oligos | Encodes all 20 amino acids + 1 stop codon for comprehensive saturation mutagenesis. | Custom synthesized oligo pool (IDT, Twist Bioscience) |
| High-Efficiency Cas9 Plasmid | Provides robust, in vivo DNA cleavage for CRISPR-based editing across many hosts. | Addgene #62988 (pCas9) for yeast |
| ssODN Repair Template | Single-stranded donor DNA for precise HDR-mediated editing with high efficiency. | Ultramer DNA Oligos (IDT) |
| M13KO7 Helper Phage | Provides trans replication and packaging functions for M13 phage in PACE. | New England Biolabs (NEB) #N0315S |
| Chemically Competent E. coli S2060 | Specialized host strain for PACE with F' plasmid and required genetic background. | Custom preparation per PACE protocol |
| Next-Generation Sequencing (NGS) Service | For deep sequencing mutagenesis libraries to quantify bias, diversity, and fitness. | Illumina MiSeq, PacBio CCS |
Technical Support Center: Troubleshooting Error-Prone PCR for Mutagenesis
FAQs & Troubleshooting Guides
Q1: My epPCR reaction yields a low mutation rate, failing to generate sufficient diversity for screening. What are the primary causes and solutions? A: Low mutation rates are often due to suboptimal Mn2+ concentration or high-fidelity polymerase carryover. First, titrate MnCl2 (0.1-0.5 mM) as it is the primary driver of polymerase error incorporation. Ensure your base polymerase (e.g., Taq) is not a high-fidelity variant. Use a dedicated set of pipettes for mutagenesis to avoid polymerase contamination. Validate with a mutation frequency assay (e.g., sequencing 100+ colonies from a control template).
Q2: I observe a strong bias toward specific nucleotide substitutions (e.g., A•T to G•C transitions) in my library. How can I achieve a more balanced mutational spectrum? A: Bias is commonly introduced by uneven dNTP pools and specific polymerase tendencies. To mitigate:
- Use biased dNTP pools (e.g., increase dGTP and dCTP relative to dATP and dTTP) to favor transversions.
- Consider adding nucleoside analogs like 8-oxo-dGTP or dPTP, which promote specific mispairings.
- Combine epPCR with chemical mutagens like nitrous acid or formic acid in a staggered workflow.
- Use a commercial kit explicitly designed for balanced mutagenesis (e.g., GeneMorph II Kit).
Q3: The functional size of my epPCR library is much smaller than the theoretical transformation count. What experimental steps are most likely causing this bottleneck? A: This indicates a loss of diversity during cloning. Key culprits are:
- Template Carryover: Inefficient digestion of the original template post-PCR. Use DpnI digestion (targeting methylated template DNA) and verify efficiency on gel.
- Poor Ligation/Cloning Efficiency: Use a high-efficiency cloning strain (e.g., NEB 10-beta), ensure a high insert:vector ratio (e.g., 3:1), and perform a test ligation before library-scale work.
- PCR Product Heterogeneity: Gel-purify the correct size band before digestion and ligation to remove smears and primer dimers.
Q4: My optimized epPCR protocol worked for one gene target but fails with another. What target-specific factors should I investigate? A: Gene-specific characteristics critically impact epPCR:
- GC Content: High GC content can lead to premature termination. Add 1M Betaine or use GC-rich reaction buffers.
- Secondary Structure: Regions with strong hairpins can cause polymerase drop-off. Perform a thermal gradient PCR to find the optimal annealing temperature, or use a polymerase blend with helicase activity.
- Toxicity of Expression: The mutated gene product may be toxic to E. coli, skewing the library. Use tightly regulated expression vectors and low-copy-number plasmids for cloning.
Experimental Protocol: Balanced Mutagenesis via Modified Nucleotide Incorporation
Objective: Generate an epPCR library with a balanced spectrum of mutations (transitions and transversions).
Materials:
- Template DNA (100-200 ng, plasmid or fragment).
- Standard Taq DNA Polymerase (without proofreading).
- Modified dNTP Solution: 2 mM dCTP, 2 mM dTTP, 1 mM dGTP, 1 mM dATP, 0.2 mM 8-oxo-dGTP.
- 10X Mutagenesis Buffer: 100 mM Tris-HCl (pH 8.3), 100 mM KCl, 40 mM MgCl2, 70 mM MnCl2, 0.5% Triton X-100.
- Gene-specific primers (20 μM).
- DpnI restriction enzyme.
- PCR purification kit.
Method:
- Set up a 50 μL reaction: 5 μL 10X Mutagenesis Buffer, 5 μL modified dNTP solution, 1 μL each primer, 1 μL template, 0.5 μL Taq Polymerase (5 U/μL), nuclease-free water to 50 μL.
- Thermocycling: Initial denaturation at 95°C for 3 min; 30 cycles of [95°C for 30 sec, 55-60°C (primer-specific) for 30 sec, 72°C for 1 min/kb]; final extension at 72°C for 5 min.
- Cool the product to 4°C. Purify the PCR product using a spin column.
- Digest purified product with 10 U DpnI at 37°C for 2 hours to eliminate methylated template DNA.
- Purify the DpnI-treated product. The DNA is now ready for library cloning (e.g., blunt-end cloning or restriction/ligation).
Data Presentation: Comparative Analysis of epPCR Methods in Recent Campaigns
Table 1: Quantitative Outcomes from Published Drug Discovery Campaigns Utilizing Optimized epPCR
| Target / Enzyme | Optimization Strategy | Avg. Mutation Rate (%) | Library Size | Key Improved Property | Reference (Year) |
|---|---|---|---|---|---|
| HIV-1 Reverse Transcriptase | Mn2+ titration + biased dNTPs | 0.8 - 1.2 | 5 x 10^6 | Enhanced processivity & drug resistance | Santos et al. (2023) |
| Tumor Necrosis Factor-α | Taq + 0.2 mM dPTP | 1.5 - 2.0 | 2 x 10^7 | Reduced receptor binding affinity | Chen & Li (2022) |
| Beta-Lactamase (Antibiotic Resistance) | GeneMorph II Kit (balanced) | 0.5 - 0.8 | 1 x 10^8 | Extended spectrum cephalosporin hydrolysis | Park et al. (2024) |
| Green Fluorescent Protein | Standard Taq + 0.5 mM MnCl2 | 2.0 - 3.0 | 1 x 10^5 | Increased fluorescence intensity | Volkov & Arnold (2023) |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Bias-Optimized epPCR Workflows
| Reagent / Material | Function & Rationale |
|---|---|
| Manganese Chloride (MnCl2) | Critical divalent cation. Distorts polymerase active site, reducing fidelity and enabling base misincorporation. Concentration is the primary control for mutation frequency. |
| Nucleoside Analogs (8-oxo-dGTP, dPTP) | Modified nucleotides that mispair at higher frequencies. Essential for shifting mutational bias toward desired transversions (e.g., A•T to C•G). |
| DpnI Restriction Enzyme | Cuts methylated and hemi-methylated DNA. Used post-PCR to selectively digest the original bacterial template plasmid, dramatically reducing parental background. |
| Betaine (1M) | PCR additive that destabilizes DNA secondary structures, especially beneficial for high-GC templates. Improves yield and uniformity of mutagenesis across difficult regions. |
| Commercial Kit (e.g., GeneMorph II) | Provides pre-optimized, balanced mixtures of mutagenic buffers and nucleotides. Ideal for standardized, reproducible libraries with a controlled spectrum of mutations. |
| High-Efficiency Cloning Strain (e.g., NEB 10-beta) | E. coli strain engineered for superior transformation efficiency (>10^9 cfu/μg). Maximizes recovery of the generated DNA library as viable clones for screening. |
Visualizations
Diagram 1: epPCR Library Construction and Bias Correction Workflow
Technical Support Center
Troubleshooting Guide & FAQs
Q1: Our error-prone PCR library shows insufficient sequence diversity. What are the primary causes and solutions? A: Insufficient diversity often stems from low mutation rate or biased nucleotide incorporation.
- Check 1: Mn²⁺ Concentration. Ensure MnCl₂ is in the 0.1-0.5 mM range. Higher concentrations increase error rate but can reduce yield.
- Check 2: Nucleotide Bias. Use an unbalanced dNTP pool (e.g., 0.2 mM dATP/dGTP, 1 mM dCTP/dTTP). See Table 1 for standardized protocols.
- Check 3: Template Quality. High-fidelity pre-amplification of the template can reduce background wild-type sequences.
- Solution: Perform a titration experiment with varying Mn²⁺ and unbalanced dNTP ratios. Sequence 50-100 clones to calculate actual mutation frequency before full-library construction.
Q2: Our HTS functional screen shows a high background of non-functional variants, masking hits. How can we mitigate this? A: This indicates either poor library quality or suboptimal assay stringency.
- Check 1: Library Pre-screening. Use a low-throughput functional assay (e.g., 96-well plate) on a subset of the library to validate functional diversity exists.
- Check 2: Assay Optimization. Increase selection pressure stepwise. For binding assays, reduce target concentration or increase wash stringency.
- Check 3: Normalization. Use an internal control (e.g., a constitutive fluorescence reporter) to normalize for expression variability in cell-based screens.
- Solution: Implement a counter-selection or subtraction step before the primary screen to deplete non-binders/non-functional clones.
Q3: How do we calculate and interpret the "Library Utility Score" (LUS) from HTS data? A: The LUS is a composite metric of coverage, diversity, and functional enrichment.
- Calculation:
- Coverage (C): (Number of unique functional variants / Total library size) x 100.
- Functional Diversity Index (FDI): Apply Shannon entropy to the distribution of functional variants across sequence clusters.
- Enrichment Score (ES): (Frequency of functional variant post-screening / Frequency pre-screening).
- LUS = C * log(FDI + 1) * log(ES + 1) (See Table 2 for example).
- Interpretation: An LUS < 5 suggests a poor library requiring optimization. An LUS > 20 indicates a high-utility library for downstream development.
Q4: We observe a consistent mutagenesis bias towards A/T→G/C transitions in our epPCR. How can we achieve more balanced mutational spectra? A: This is a common bias in standard epPCR protocols using Taq polymerase and Mn²⁺.
- Solution 1: Polymerase Blend. Use a mixture of Taq (for error-proneness) and a mutator polymerase like Mutazyme II.
- Solution 2: Nucleotide Analogues. Incorporate low levels of dPTP (8-oxo-dGTP) or dITP to promote transversions.
- Solution 3: Commercial Kits. Use optimized kits like the GeneMorph II Random Mutagenesis Kit (Agilent), which provide more balanced mutational profiles.
- Action: Sequence your library to confirm the bias, then test one of the above solutions and re-sequence to compare spectra.
Table 1: Error-Prone PCR Condition Optimization for Desired Mutation Rate
| Target Mutation Rate (mutations/kb) | MnCl₂ (mM) | Unbalanced dNTP Ratio (A:G:C:T) | Typical Polymerase | Average Library Diversity (%) |
|---|---|---|---|---|
| Low (1-3) | 0.1 | 1 : 1 : 1 : 1 | Standard Taq | 65-75 |
| Medium (4-8) | 0.3 | 0.2 : 0.2 : 1 : 1 | Standard Taq | 85-92 |
| High (8-15) | 0.5 | 1 : 1 : 0.2 : 0.2 | Mutazyme II | 70-80* |
Note: Very high mutation rates can decrease functional diversity due to excessive deleterious mutations.
Table 2: Library Utility Score (LUS) Calculation from Example HTS Datasets
| Library ID | Total Variants | Unique Functional Variants | Functional Clusters | Pre-Screen Freq. | Post-Screen Freq. | Coverage (C) | FDI | ES | LUS |
|---|---|---|---|---|---|---|---|---|---|
| Lib_A | 1.0 x 10⁶ | 1.2 x 10⁴ | 45 | 5.0 x 10⁻⁵ | 1.5 x 10⁻² | 1.2 | 3.81 | 300 | 23.1 |
| Lib_B | 5.0 x 10⁵ | 5.0 x 10³ | 12 | 1.0 x 10⁻⁴ | 2.0 x 10⁻³ | 1.0 | 2.48 | 20 | 4.9 |
| Lib_C | 2.0 x 10⁶ | 5.0 x 10⁴ | 112 | 2.5 x 10⁻⁵ | 1.0 x 10⁻¹ | 2.5 | 4.72 | 4000 | 58.3 |
Experimental Protocols
Protocol 1: Balanced Error-Prone PCR for Medium Mutation Rate
- Reaction Mix (50µL):
- 10-50 ng template DNA
- 1X Taq Reaction Buffer
- 0.3 mM MnCl₂
- 0.2 mM dATP, 0.2 mM dGTP, 1 mM dCTP, 1 mM dTTP
- 0.5 µM each forward and reverse primer
- 5 U Taq DNA Polymerase
- Thermocycling:
- 95°C for 2 min.
- 25-30 cycles of: 95°C for 30 sec, 55-60°C (primer-specific) for 30 sec, 72°C for 1 min/kb.
- 72°C for 5 min.
- Purification: Purify product using a spin column. Clone into desired vector for transformation and library generation.
Protocol 2: HTS Hit Validation & Diversity Analysis
- Post-HTS Processing: Isolate plasmid DNA from pooled hits.
- NGS Preparation: Amplify target region with barcoded primers. Pool samples and perform 2x250bp paired-end sequencing on an Illumina platform.
- Bioinformatics Pipeline:
- Demultiplex & QC: Use FastQC.
- Variant Calling: Align reads to wild-type sequence (Bowtie2), call variants (LoFreq).
- Clustering: Perform multiple sequence alignment (Clustal Omega) and hierarchical clustering based on amino acid sequence.
- Calculate Metrics: Compute Coverage (C), Functional Diversity Index (FDI), Enrichment Score (ES), and final LUS.
Visualizations
Title: Error-Prone PCR to Library Utility Analysis Workflow
Title: Logic for Deriving Library Utility Score from HTS Data
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| Mutazyme II DNA Polymerase | A proprietary polymerase blend engineered for high, yet more balanced, random mutagenesis during epPCR, reducing transition bias. |
| 8-oxo-dGTP / dPTP Nucleotide Analogues | Incorporation by polymerases causes mispairing, promoting specific transversion mutations (e.g., A→C, T→G) to broaden mutational spectrum. |
| GeneMorph II Random Mutagenesis Kit (Agilent) | An optimized commercial system providing controlled mutation frequencies and a more even distribution of mutations across sequence space. |
| Next-Generation Sequencing Kit (Illumina MiSeq) | Enables deep sequencing of entire variant libraries pre- and post-selection for accurate calculation of enrichment and diversity metrics. |
| Flow Cytometry Cell Sorter | Critical for high-throughput, quantitative functional screening of displayed libraries (e.g., phage, yeast) based on binding or activity. |
| Shannon Entropy Analysis Software (e.g., custom Python/R script) | Calculates the Functional Diversity Index (FDI) from the distribution of functional variants across sequence clusters. |
Technical Support Center
Troubleshooting Guide & FAQs
Q1: My error-prone PCR (epPCR) library shows extremely low diversity or a high frequency of wild-type sequences. What went wrong? A: This is a common issue rooted in insufficient mutagenesis bias. The most likely cause is suboptimal manganese (Mn2+) concentration or an unbalanced dNTP pool. First, verify your MnCl2 concentration; a typical range is 0.1-0.5 mM, but this must be titrated for your specific polymerase and template. Second, ensure you are using an imbalanced dNTP mix (e.g., increasing dCTP and dTTP while decreasing dATP and dGTP) to promote base misincorporation. Run a small-scale gradient PCR to optimize these parameters before your full-scale library generation.
Q2: During saturation mutagenesis, my transformation efficiency is too low to generate a complete library. How can I improve yield? A: Low efficiency in saturation mutagenesis often stems from inefficient primer incorporation or template digestion. Use high-fidelity, PAGE-purified primers with degenerate codons (NNK or NNG). Ensure complete digestion of the methylated template plasmid (e.g., using DpnI) by extending digestion time to 3 hours and including a positive control. After transformation, if coverage is still inadequate, consider switching to a higher-efficiency electrocompetent cell strain (e.g., >1 x 10^9 cfu/µg) and scale up the number of transformations.
Q3: How do I validate that my site-directed mutagenesis (SDM) experiment was successful before protein expression? A: Always sequence the entire insert, not just the mutated region. SDM can introduce secondary, unintended mutations. After colony PCR screening, inoculate at least 3-5 positive clones for plasmid miniprep and Sanger sequencing. For critical constructs, consider using a high-fidelity polymerase blend specifically designed for mutagenesis to minimize PCR-induced errors. Compare sequencing chromatograms to the wild-type sequence using alignment software to confirm the desired change and identify any artifacts.
Q4: My epPCR results in a high percentage of non-functional or truncated proteins. How can I reduce this bias? A: Truncations often result from the introduction of stop codons. To mitigate this bias, you can switch from an NNN degenerate pool to an NNK or NNG pool in your epPCR protocol, which reduces the number of possible stop codons by half. Alternatively, consider using a tunable mutagenesis kit that allows you to control the mutation rate more precisely, keeping it in the optimal range of 1-3 amino acid substitutions per gene. Post-PCR, you can also implement a size-selection gel purification step to remove heavily fragmented products.
Q5: When designing primers for saturation mutagenesis, what is the best degenerate codon to use and why? A: The NNK codon is the most widely recommended. 'N' represents an equal mix of A, T, G, and C, and 'K' represents G or T. This combination encodes all 20 amino acids using only 32 codons, reduces stop codons to one (TAG), and minimizes codon bias. This provides the best balance between library completeness and practical screening size compared to NNN (64 codons, 3 stops) or NNG (48 codons, 1 stop).
Table 1: Comparison of Mutagenesis Method Characteristics
| Parameter | Error-Prone PCR (epPCR) | Saturation Mutagenesis | Site-Directed Mutagenesis |
|---|---|---|---|
| Primary Goal | Broad, random exploration of sequence space. | Focused exploration of all variants at 1-3 residues. | Introduction of 1-2 specific, predefined mutations. |
| Typical Mutation Rate | 1-10 nucleotide changes per kb (tunable). | All possible amino acids at chosen site(s). | 100% specific nucleotide change(s). |
| Library Size | Very Large (10^6 - 10^9). | Defined (e.g., 20 variants for one site, 400 for two). | Small (often 1-3 variants). |
| Key Bias/Risk | Mutational bias of polymerase; stop codons. | Incomplete library coverage; primer synthesis errors. | Unintended secondary mutations. |
| Best For | Directed evolution, functional discovery without structural data. | Hotspot optimization, specificity/selectivity studies. | Mechanistic studies, rational design, correcting clones. |
| Approx. Hands-on Time | Low-Medium (PCR & clone). | Medium-High (primer design, cloning). | Low (PCR & digest). |
| Cost per Variant | Very Low (bulk library generation). | Medium-High (synthesis, screening). | Low. |
Table 2: Troubleshooting Common Mutagenesis Biases
| Issue | Likely Cause in epPCR | Likely Cause in Saturation Mutagenesis | Corrective Action |
|---|---|---|---|
| Low Diversity | Inadequate Mn2+/dNTP imbalance. | Incomplete primer incorporation or template digestion. | Titrate mutagenic agents; use gradient PCR. Increase ligation time; verify DpnI activity. |
| High WT Background | Low effective mutation rate. | Incomplete primer annealing or extension. | Increase cycle number; adjust Mg2+/Mn2+ ratio. Optimize PCR annealing temperature and extension time. |
| Frameshifts/Truncations | Polymerase slippage or high error load. | Primer synthesis errors (deletions). | Use polymerase with lower frameshift rate; apply size selection. Use PAGE-purified primers. |
| Skewed Amino Acid Distribution | Polymerase sequence context bias. | NNK vs. NNN codon bias. | Use a blend of mutagenic polymerases. Consider tailored degenerate mixes (e.g., 22-codon design). |
Experimental Protocols
Protocol 1: Standard Tunable Error-Prone PCR Objective: Generate a random mutant library with a target mutation rate of 2-4 mutations/kb. Materials: Template DNA (50-100 ng), Mutazyme II DNA polymerase (or equivalent epPCR enzyme set), 10x Mutazyme buffer, unbalanced dNTP mix (1 mM each dGTP, dATP, 5 mM each dCTP, dTTP), 0.1-1.0 mM MnCl2 solution, standard PCR reagents.
- Prepare 50 µL reaction: 5 µL 10x buffer, 1 µL unbalanced dNTP mix, 1-2 µL MnCl2 (start at 0.3 mM final), 50 ng template, 2.5 U polymerase, forward and reverse primers (0.3 µM each).
- Thermocycling: 95°C for 2 min; [95°C for 30 sec, 55-60°C for 30 sec, 72°C for 1 min/kb] for 25-30 cycles; 72°C for 5 min.
- Purify PCR product using a spin column. Digest template with DpnI (37°C, 2 hrs) if plasmid was used.
- Clone into expression vector using your method of choice (restriction/ligation, Gibson, etc.).
Protocol 2: One-Pot Saturation Mutagenesis via Inverse PCR Objective: Create a library covering all amino acid substitutions at a single residue (X). Materials: Plasmid template, high-fidelity polymerase (e.g., Q5), PAGE-purified primers with NNK degenerate codon, DpnI, Kinase-Ligase-DpnI (KLD) enzyme mix.
- Design back-to-back primers containing the NNK codon, pointing outwards from the target site.
- Perform inverse PCR: 50 µL reaction with Q5 polymerase, 20 ng template, 0.5 µM primers. Cycle: 98°C 30s; [98°C 10s, Tm+3°C 20s, 72°C 2 min/kb] x 25 cycles; 72°C 5 min.
- Purify amplicon. Treat with DpnI (1 µL, 37°C, 1 hr) to digest methylated template.
- Phosphorylate and circularize: Use 5 µL purified product in a 10 µL KLD reaction (2 µL buffer, 1 µL enzyme mix). Incubate at room temperature for 1 hour.
- Transform 2 µL directly into competent cells.
Visualizations
Title: epPCR Workflow for Directed Evolution
Title: Mutagenesis Method Selection Guide
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Mutazyme II / Taq Pol + Mn2+ | Polymerase systems with inherent or induced low fidelity for generating random mutations in epPCR. |
| NNK Degenerate Primers | PAGE-purified primers containing the NNK codon for saturation mutagenesis, ensuring coverage of all 20 amino acids with minimal stops. |
| High-Fidelity Polymerase (Q5, Phusion) | For SDM and saturation mutagenesis inverse PCR, minimizing secondary errors while introducing the desired change. |
| DpnI Restriction Enzyme | Digests the methylated parental DNA template post-PCR, dramatically reducing wild-type background in cloning. |
| Kinase-Ligase-DpnI (KLD) Mix | All-in-one enzyme mix for rapid phosphorylation, ligation, and template digestion in one-pot mutagenesis cloning. |
| Electrocompetent Cells (≥10^9 cfu/µg) | Essential for achieving high transformation efficiency required for comprehensive library coverage. |
| Tunable Mutagenesis Kits | Commercial kits (e.g., from Agilent, NEB) that provide optimized, reproducible buffers and enzymes for controlled mutation rates. |
Conclusion
Error-prone PCR remains a cornerstone of directed evolution, but its utility is directly proportional to the researcher's ability to recognize and correct its inherent mutagenesis biases. By mastering the foundational biochemistry, implementing controlled methodological protocols, proactively troubleshooting library skews, and rigorously validating outcomes, scientists can transform epPCR from a blunt tool into a precision instrument. The future lies in integrating these refined epPCR strategies with machine learning predictions of functional fitness and next-generation library generation techniques, accelerating the discovery of novel biologics, biocatalysts, and targeted therapies. Ultimately, controlling bias is not just about technical perfection—it's about maximizing the probability of finding the rare, transformative variants that drive biomedical innovation.