Unlocking Cellular Drug Targets: A Comprehensive Guide to System-Wide Substrate Identification with SIESTA Thermal Profiling
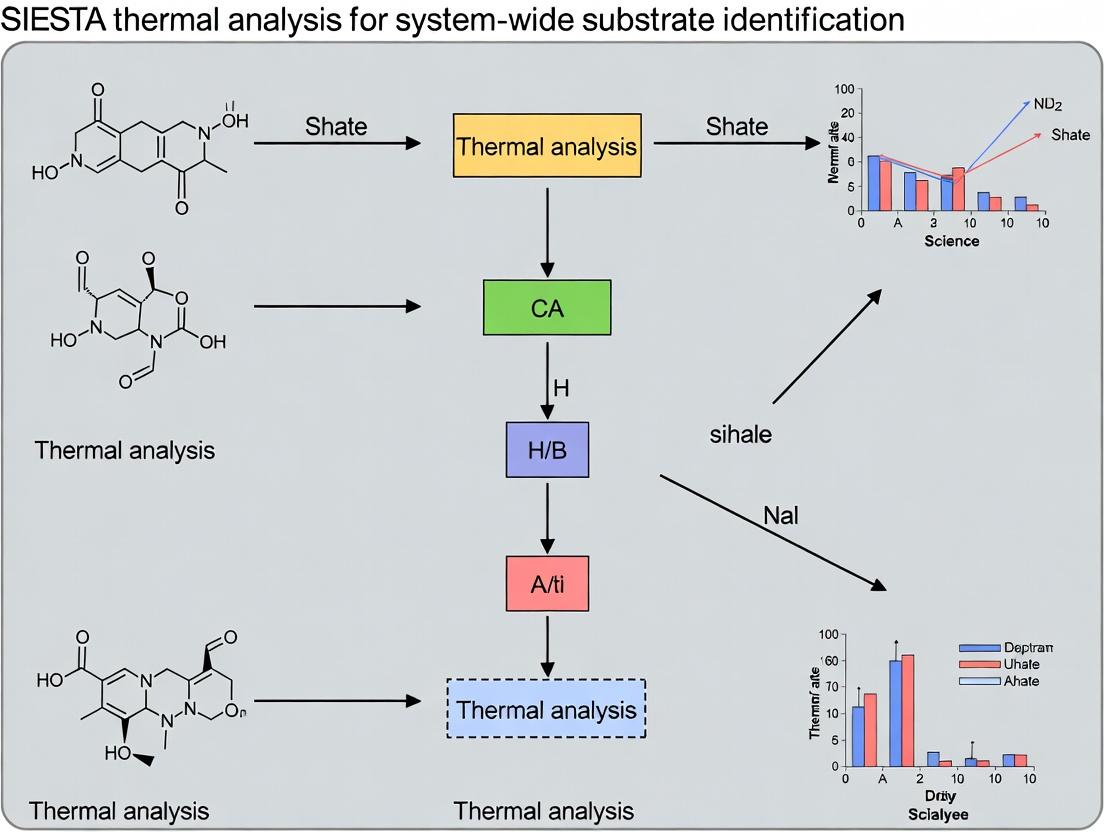
This article provides a complete framework for researchers and drug development professionals on implementing and applying the Stability of Proteins from Rates of Oxidation (SIESTA) thermal profiling technique.
Unlocking Cellular Drug Targets: A Comprehensive Guide to System-Wide Substrate Identification with SIESTA Thermal Profiling
Abstract
This article provides a complete framework for researchers and drug development professionals on implementing and applying the Stability of Proteins from Rates of Oxidation (SIESTA) thermal profiling technique. It covers the foundational principles of thermal proteome profiling (TPP) and SIESTA's unique chemoproteomic approach for system-wide, unbiased identification of drug-protein interactions and metabolic enzyme substrates. The guide details methodological workflows from experimental setup to data analysis, addresses common troubleshooting and optimization challenges, and validates SIESTA against other techniques like CETSA and LiP-MS. The conclusion synthesizes key takeaways and discusses future implications for target deconvolution, polypharmacology, and clinical biomarker discovery in biomedical research.
What is SIESTA Thermal Analysis? Foundational Principles for Unbiased Target Discovery
Thermal Proteome Profiling (TPP) is a mass spectrometry-based, proteome-wide implementation of the Cellular Thermal Shift Assay (CETSA). It quantitatively measures protein thermal stability changes across the proteome in response to small molecules, environmental perturbations, or protein interactions. The method was pioneered by Savitski et al. (2014) and has since evolved into a cornerstone technique for system-wide target deconvolution, mechanism-of-action studies, and biomarker discovery.
The broader thesis of SIESTA (System-wide Identification of Enzyme Substrates by Thermal Analysis) positions TPP as a foundational tool. SIESTA extends the principle by applying thermal profiling not just to drug binding, but to enzymatic activity, aiming to map substrate networks by detecting stability changes in enzymes and their interacting partners upon substrate conversion.
Core Principles and Workflows
The fundamental principle is that ligand binding or post-translational modification often alters a protein's thermal stability, shifting its denaturation curve. TPP measures this shift by subjecting living cells or cell lysates to a gradient of temperatures, followed by fractionation of soluble (non-denatured) proteins, tryptic digestion, and quantitative tandem mass tag (TMT)-based LC-MS/MS analysis.
Key Experimental Designs:
- Temperature Range (TR): A single sample is heated to multiple temperatures.
- Compound Concentration Range (CCR): Samples treated with a compound at multiple concentrations are heated at one or two temperatures.
- 2D-TPP: Combines both temperature and compound concentration gradients for higher confidence.
Application Notes
TPP applications in drug discovery and systems biology include:
- Target Identification: Unbiased discovery of drug targets and off-targets.
- Mechanism of Action: Classifying compounds by their thermal profiles.
- Epigenetic Profiling: Mapping interactions of readers, writers, and erasers with chromatin.
- Metabolic Studies (SIESTA context): Identifying enzyme-substrate pairs by detecting thermal shifts upon metabolic perturbation.
- Biomarker Discovery: Identifying thermally stable protein signatures in disease.
Table 1: Comparison of TPP Methodologies
| Method | Key Principle | Throughput | Key Readout | Primary Application |
|---|---|---|---|---|
| CETSA (Classical) | Protein aggregation detection via WB/HRM. | Low (single proteins) | Melting point (Tm) shift. | Validation of specific target engagement. |
| TPP (TR/CCR) | Proteome-wide solubility via MS. | High (10,000+ proteins) | Apparent melting curves (Tmelt). | System-wide target deconvolution. |
| 2D-TPP | Combined temp & conc. gradient. | Medium-High | Dose-dependent stability curves. | High-confidence target ranking & affinity estimation. |
| SIESTA | Thermal profiling of enzyme activity. | High | Stability changes upon substrate conversion. | System-wide mapping of enzyme-substrate relationships. |
Table 2: Representative TPP Studies and Outputs
| Study Focus | System | Key Finding (Number of Hits) | Reference (Example) |
|---|---|---|---|
| Kinase Inhibitor | K562 cells | Confirmed known targets & identified novel off-targets of kinase drugs (e.g., ~10 proteins with ΔTm >2°C for staurosporine). | Savitski et al., Science, 2014. |
| Epigenetic Probe | MOLT-4 cells | Identified BET family bromodomains as targets of JQ1, and distinguished its profile from I-BET151. | Franken et al., Nat Protoc, 2015. |
| Metabolic Enzyme (SIESTA) | Cell Lysate | Mapping of ADP-ribosyltransferase substrates by detecting thermally stabilized complexes (>100 substrates identified). | Larsen et al., Cell, 2018. |
| Next-Gen (PISA) | In vitro Proteome | Multiplexed direct measurement of protein abundance & thermal stability in one pot. | Childs et al., Nat Biotechnol, 2019. |
Detailed Experimental Protocols
Protocol 1: Basic TPP Workflow (Temperature Range in Intact Cells)
Objective: To identify proteins with altered thermal stability upon drug treatment.
Materials: See "Scientist's Toolkit" below. Procedure:
- Cell Culture & Treatment: Grow HeLa cells to 80% confluency. Treat with compound of interest (e.g., 10 µM) or vehicle (DMSO) for 1 hour.
- Heating: Harvest cells, wash with PBS. Aliquot cell suspensions into PCR tubes (~1 million cells/tube). Heat aliquots across a temperature gradient (e.g., 37°C to 67°C in 3°C increments, 10 temperatures) for 3 minutes in a thermal cycler.
- Lysis & Soluble Protein Harvest: Immediately place tubes on ice. Add cold lysis buffer (with protease inhibitors, nuclease). Freeze-thaw cycle (liquid N₂/37°C) 3x. Centrifuge at 20,000 x g, 4°C for 20 min.
- Protein Digestion: Transfer supernatant (soluble fraction) to new tubes. Quantify protein. Reduce (DTT), alkylate (IAM), and digest with trypsin (overnight, 37°C).
- TMT Labeling: Desalt peptides. Label each temperature point from treated and control samples with a unique isobaric TMT 10-plex or 11-plex reagent (1hr, RT). Quench reaction with hydroxylamine. Pool all labeled samples.
- LC-MS/MS Analysis: Fractionate pooled sample by basic pH reverse-phase HPLC. Analyze each fraction by LC-MS/MS on an Orbitrap instrument (e.g., Eclipse Tribrid) with MS3-based SPS method to reduce ratio compression.
- Data Analysis: Process raw files with IsobarQuant and TPP software suite (or
NPARCR package). Fit dose-response curves per protein to calculate apparent melting temperature (Tm). Significant hits are defined by ΔTm > 2°C and p-value < 0.05 (FDR-corrected).
Protocol 2: SIESTA-Informed Substrate Identification Protocol
Objective: To identify system-wide substrates of an enzyme by detecting thermal co-stability. Procedure:
- Lysate Preparation: Prepare clarified lysate from relevant tissue or cell line in appropriate activity buffer.
- Enzymatic Reaction Setup: Incubate lysate with:
- Condition A: Active enzyme + necessary cofactors (e.g., NAD⁺).
- Condition B: Inactive enzyme mutant (or heat-inactivated) + cofactors.
- Condition C: Active enzyme + cofactors + a known competitive inhibitor. Incubate for time T at 37°C to allow reaction.
- Thermal Profiling: Immediately subject reaction mixtures to the TPP (TR) protocol (Steps 2-7 from Protocol 1).
- Data Analysis: Identify proteins whose thermal stability is significantly increased in Condition A compared to both Conditions B and C. These proteins are potential direct substrates or members of stabilized complexes.
Visualization of Workflows and Pathways
TPP Experimental Workflow
Diagram 1: TPP Experimental Workflow
SIESTA Conceptual Framework
Diagram 2: SIESTA Framework for Substrate ID
CETSA to Next-Gen TPP Evolution
Diagram 3: Evolution from CETSA to Next-Gen TPP
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for TPP
| Item | Function in TPP/SIESTA | Key Consideration |
|---|---|---|
| Tandem Mass Tags (TMTpro 16-plex) | Isobaric labeling reagents for multiplexed quantification of peptides across up to 16 samples (temperatures/conditions). | Enables high-throughput profiling. MS3 methods required for accurate quantification. |
| Lysis Buffer (NP-40 based) | Gentle, non-denaturing detergent to lyse cells after heating, preserving native protein complexes. | Must be optimized to minimize background aggregation. Protease/nuclease inhibitors are essential. |
| Trypsin (Sequencing Grade) | Protease for digesting soluble proteins into peptides for MS analysis. | High purity and activity ensure complete, reproducible digestion. |
| Thermostable Enzymes (for SIESTA) | Active wild-type and catalytically dead mutant enzymes for comparative activity perturbation. | Critical control for distinguishing binding from catalysis-induced stabilization. |
| Phosphate-Buffered Saline (PBS) | Iso-osmotic suspension buffer for heating intact cells. | Must be free of stabilizing agents like BSA that would confound the assay. |
| High-pH Reverse-Phase HPLC Columns | For fractionating complex peptide mixtures pre-MS to increase proteome depth. | Reduces peptide co-elution and increases protein identifications (>7000 typical). |
| Data Analysis Suite (IsobarQuant/TPP) | Open-source software for processing TMT raw data, curve fitting, and calculating ΔTm. | Requires computational expertise. Alternative: commercial software like Thermo Fisher Proteome Discoverer with TPP plugin. |
| Cell Permeabilizers (e.g., Digitonin) | For studying membrane-impermeable compounds or metabolites in a cellular context (limited permeability CETSA). | Allows controlled access to intracellular targets while maintaining cellular architecture. |
SIESTA (Stability of proteins from Rates of Oxidation) is a high-throughput thermal profiling method that quantifies protein stability on a proteome-wide scale by measuring the rate of methionine oxidation by hydrogen peroxide as a function of temperature. This application note details the protocols for implementing SIESTA within a system-wide substrate identification research framework, providing researchers with a robust tool for identifying drug targets, mapping ligand-induced stabilization, and probing protein-ligand interactions.
The core principle of SIESTA is that the rate of methionine oxidation by H₂O₂ is exquisitely sensitive to protein conformational stability. In its native state, methionine residues are buried and protected. As temperature increases, protein unfolding exposes these residues, leading to a sharp increase in oxidation rate. The temperature at which this transition occurs (Tox) is analogous to the melting temperature (Tm) and serves as a quantitative metric of protein thermal stability. By combining this with tandem mass spectrometry (MS/MS), stability profiles for thousands of proteins can be generated in parallel.
Table 1: Key Metrics and Performance Characteristics of SIESTA
| Parameter | Typical Value/Range | Significance |
|---|---|---|
| Temperature Range | 37°C - 67°C (increments of 2-3°C) | Covers unfolding transitions for most cellular proteins. |
| H₂O₂ Concentration | 0.1% - 0.3% (v/v) | Optimized for sufficient oxidation signal without excessive background. |
| Incubation Time | 3 minutes | Standardized reaction window for oxidation. |
| Key Readout (Tox) | Protein-specific (e.g., 45°C - 60°C) | The inflection point in the oxidation rate curve; indicates stability. |
| ΔTox Significance | > 2°C considered significant | Shift induced by ligand binding, mutations, or post-translational modifications. |
| Proteome Coverage | > 5,000 proteins per experiment | Enables system-wide analysis. |
| Replicate Correlation (R²) | > 0.95 | High technical reproducibility. |
Table 2: Example SIESTA Data for Model Protein-Ligand Interaction
| Protein (Target) | Condition | Mean Tox (°C) | Std. Dev. | ΔTox vs. DMSO |
|---|---|---|---|---|
| Kinase ABC | DMSO Control | 48.2 | ± 0.5 | - |
| Kinase ABC | 10 µM Inhibitor X | 53.7 | ± 0.4 | +5.5 |
| Protein XYZ | DMSO Control | 51.8 | ± 0.6 | - |
| Protein XYZ | 10 µM Inhibitor X | 51.5 | ± 0.5 | -0.3 |
Detailed Experimental Protocols
Protocol 3.1: SIESTA Thermal Profiling for Lysate Samples
Objective: To generate thermal stability profiles for proteins in a complex cell lysate.
Materials: See "Scientist's Toolkit" (Section 5). Procedure:
- Lysate Preparation: Lyse cells in SIESTA lysis buffer (e.g., PBS with 1% NP-40, protease inhibitors). Clarify by centrifugation (16,000 x g, 15 min, 4°C). Determine protein concentration.
- Aliquoting: Dispense 50 µL of lysate (1-2 mg/mL total protein) into PCR tubes or a 96-well PCR plate.
- Thermal Challenge: Using a thermal cycler, incubate replicate aliquots at a series of temperatures (e.g., 37, 40, 43, 46, 49, 52, 55, 58, 61, 64, 67°C) for 3 minutes.
- Oxidation Reaction: Immediately add 5 µL of 3% H₂O₂ solution (freshly diluted) to each heated sample to achieve a final concentration of ~0.3%. Incubate at room temperature for 2 minutes.
- Quenching: Add 5 µL of 50 mM methionine solution to quench the H₂O₂. Place samples on ice.
- Reduction, Alkylation, and Digestion: Add DTT to 5 mM (10 min, 56°C), then iodoacetamide to 15 mM (30 min, dark, RT). Digest proteins with trypsin/Lys-C overnight at 37°C.
- Peptide Cleanup: Desalt peptides using C18 stage tips. Dry in a vacuum concentrator.
- LC-MS/MS Analysis: Resuspend peptides in LC loading buffer. Analyze by liquid chromatography coupled to a high-resolution tandem mass spectrometer using a standard data-dependent acquisition (DDA) method.
Protocol 3.2: Data Processing and ToxCalculation
Objective: To calculate oxidation rates and Tox values from raw MS data.
Procedure:
- Database Search: Process raw files with a search engine (e.g., MaxQuant, Spectronaut). Search against the appropriate proteome database. Include 'methionine oxidation to methionine-sulfoxide' (+15.9949 Da) as a variable modification.
- Peptide Intensity Extraction: Extract the intensity of each methionine-containing peptide in its oxidized and non-oxidized forms across all temperature points.
- Oxidation Rate Calculation: For each peptide at each temperature (T), calculate the fractional oxidation: Fox(T) = Int(Ox) / [Int(Ox) + Int(Non-Ox)].
- Curve Fitting: Fit the Fox(T) data for each protein (using the average of its peptides) to a sigmoidal curve (e.g., Boltzmann equation) using non-linear regression.
- Tox Determination: Extract the inflection point (temperature at which the derivative is maximum) from the fitted curve. This is the Tox.
- ΔTox Analysis: Compare Tox values between treatment and control conditions. Apply statistical tests (e.g., t-test) to identify proteins with significant thermal shifts.
Visualization of Workflows and Pathways
SIESTA Experimental and Computational Workflow
Core Principle: Thermal Unfolding Drives Methionine Oxidation
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for SIESTA
| Item | Function & Specification |
|---|---|
| SIESTA Lysis Buffer | PBS, pH 7.4, supplemented with 1% NP-40 (or similar detergent) and protease/phosphatase inhibitors. Maintains native protein complexes for analysis. |
| Hydrogen Peroxide (H₂O₂) Stock | High-purity, 30% (w/w) stock. Critical: Prepare fresh dilutions (e.g., to 3%) on the day of experiment for consistent oxidation efficacy. |
| Methionine Quench Solution | 50 mM L-methionine in water. Rapidly quenches excess H₂O₂ to stop the oxidation reaction at precisely defined times. |
| MS-Grade Trypsin/Lys-C Mix | For efficient and complete protein digestion post-oxidation. Essential for reproducible peptide generation and quantification. |
| Stable Isotope-Labeled Reference Peptides | Spiked-in prior to MS analysis for absolute quantification and normalization across samples and temperature points (optional but recommended). |
| Thermal Cycler with 96-well block | Provides precise, rapid, and uniform heating of samples across the defined temperature gradient. |
| High-Resolution LC-MS System | Nanoflow liquid chromatography coupled to a Q-Exactive Orbitrap or similar high-resolution tandem mass spectrometer for accurate identification and quantification of oxidized peptides. |
| Data Analysis Software Suite | e.g., MaxQuant, Spectronaut, or custom R/Python scripts for peptide quantification, curve fitting, and statistical analysis of ΔTox. |
Classical drug discovery focuses on "target engagement"—measuring a compound's binding affinity for a specific, purified protein target. While foundational, this approach fails to capture the system-wide biochemical consequences of a drug's action within a native cellular environment. System-wide substrate identification moves beyond this singular view by globally identifying the proteomic substrates of enzymes (e.g., kinases, ligases, proteases) or the direct interacting partners of small molecules, in complex biological systems. This paradigm is essential for understanding polypharmacology, mechanism-of-action, and off-target effects.
Within this paradigm, thermal shift assays, particularly the SIESTA (Systematic Identification of Enzyme Substrates by Thermal Analysis) platform, provide a powerful, label-free methodology. SIESTA leverages the principle that protein-ligand or enzyme-substrate interactions often alter protein thermal stability. By coupling cellular thermal shift assays (CETSA) with quantitative mass spectrometry (MS), SIESTA enables the proteome-wide identification of direct drug targets and native enzyme substrates, mapping the intricate network of interactions that constitute a drug's true biological footprint.
Core Protocols & Application Notes
Protocol A: SIESTA Workflow for Kinase Substrate Identification
Objective: To identify novel, native substrates of a specific kinase in a cancer cell line (e.g., A549 cells) under stimulated vs. basal conditions.
Materials & Reagents:
- Cell culture system (A549 cells, appropriate medium)
- Kinase inhibitor (e.g., specific ATP-competitive compound) or activator
- Control vehicle (DMSO)
- Phosphate-Buffered Saline (PBS)
- Protease and phosphatase inhibitors
- Lysis buffer (e.g., RIPA or NP-40 based)
- BCA protein assay kit
- Heated lid thermal cycler or precise dry bath
- Centrifugal filters (10kDa MWCO)
- Trypsin/Lys-C mix for digestion
- Tandem Mass Tag (TMT) reagents (16-plex)
- High-pH reverse-phase fractionation kit
- LC-MS/MS system (Orbitrap-based)
Procedure:
- Cell Treatment & Harvest: Culture A549 cells to 80% confluence. Treat experimental sets with the kinase inhibitor (e.g., 1 µM, 2 hours) and control sets with equivalent DMSO. Harvest cells using trypsin, wash 3x with ice-cold PBS.
- Thermal Denaturation Series: Resuspend cell pellets in PBS + inhibitors. Aliquot equal protein amounts (from BCA assay) into PCR tubes (e.g., 10-12 aliquots per condition). Subject aliquots to a temperature gradient (e.g., 37°C to 67°C, in 3°C increments) for 3 minutes in a thermal cycler, followed by 3 minutes at room temperature.
- Soluble Protein Isolation: Lyse heated samples by freeze-thaw (liquid N2) or mechanical homogenization. Centrifuge at 20,000 x g for 20 min at 4°C to separate soluble (thermally stable) protein from aggregates.
- Protein Digestion & Multiplexing: Quantify soluble protein in each supernatant. Digest proteins using standard trypsin/Lys-C protocol. Label digested peptides from each temperature point for control and treated samples with unique TMT tags. Pool labeled samples.
- High-pH Fractionation & LC-MS/MS: Fractionate the pooled sample using high-pH reverse-phase chromatography to reduce complexity. Analyze fractions via LC-MS/MS on an Orbitrap instrument.
- Data Analysis: Process raw files using software (e.g., Proteome Discoverer, MaxQuant). Generate melting curves for every protein detected by plotting normalized TMT signal intensity (soluble fraction) vs. temperature. Calculate the temperature at which 50% of the protein is denatured (Tm). Identify substrates by significant Tm shifts (ΔTm > 2°C, p < 0.05) between inhibitor-treated and control samples, indicating a change in thermal stability due to altered phosphorylation status.
Expected Outcome: A list of proteins whose thermal stability is significantly altered upon kinase inhibition, representing direct kinase substrates or proteins in the kinase's immediate complex.
Protocol B: CETSA-MS for Drug Target Deconvolution
Objective: To identify the direct protein targets and off-targets of an uncharacterized small molecule with anti-proliferative activity.
Procedure:
- Follow Protocol A steps 1-3, comparing cells treated with the compound of interest vs. vehicle control.
- Perform isothermal dose-response (ITDR) analysis: Treat cell aliquots with a compound concentration gradient (e.g., 1 nM to 100 µM) at a single, fixed temperature (near the expected Tm of potential targets). Process and analyze via MS as above.
- Data Analysis: Generate dose-response curves from ITDR data to calculate apparent binding affinities (Kd). Combine ΔTm and dose-response data to distinguish high-affinity direct targets (which show clear, dose-dependent stabilization) from downstream, indirect effectors.
Table 1: Representative SIESTA Data Output for Kinase Inhibitor X in A549 Cells
| Protein (Gene Symbol) | Control Tm (°C) | Inhibitor Tm (°C) | ΔTm (°C) | p-value | Putative Role |
|---|---|---|---|---|---|
| MAPK1 | 52.1 ± 0.3 | 55.8 ± 0.4 | +3.7 | 1.2E-06 | Known Direct Target |
| RSK2 (RPS6KA3) | 48.5 ± 0.5 | 51.1 ± 0.4 | +2.6 | 3.5E-05 | Known Direct Target |
| FOXO1 | 49.2 ± 0.4 | 46.0 ± 0.5 | -3.2 | 8.7E-06 | Novel Substrate |
| MYC | 47.8 ± 0.3 | 45.9 ± 0.6 | -1.9 | 4.1E-03 | Downstream Effector |
| GAPDH | 58.3 ± 0.2 | 58.5 ± 0.3 | +0.2 | 0.45 | Loading Control |
Table 2: Comparison of Target Identification Techniques
| Method | Throughput | Context | Measures Direct Binding? | Label Required? | Key Limitation |
|---|---|---|---|---|---|
| SIESTA/CETSA-MS | High | Native Cellular | Yes | No | Moderate proteome coverage depth |
| Affinity Pulldown-MS | Medium | Lysate/Cellular | Yes | Yes (Tag) | High false-positive rate |
| Activity-Based Protein Profiling | Medium | Lysate/Cellular | Yes (Active site) | Yes (Probe) | Restricted to enzyme classes |
| Phosphoproteomics | High | Cellular | Indirect | No | Cannot distinguish direct substrates |
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for SIESTA Workflow
| Item | Function/Description | Example Product/Catalog |
|---|---|---|
| Cell-Permeable Kinase Inhibitor/Activator | Pharmacologically modulates target enzyme activity in live cells to perturb substrate interactions. | Selleckchem bioactive compounds; Tocris kinase modulators. |
| Tandem Mass Tag (TMT) 16/18-plex Kit | Isobaric labels for multiplexed quantitative MS, allowing simultaneous analysis of up to 18 temperature points or conditions. | Thermo Fisher Scientific, Cat# A44520 (TMT16). |
| High-pH Reverse-Phase Peptide Fractionation Kit | Reduces sample complexity prior to MS, improving proteome depth and quantification accuracy. | Pierce High pH Reversed-Phase Peptide Fractionation Kit, Cat# 84868. |
| Protease/Phosphatase Inhibitor Cocktail | Preserves native protein post-translational modification states and prevents degradation during cell lysis. | Halt Protease & Phosphatase Inhibitor Cocktail, Thermo Cat# 78440. |
| Trypsin/Lys-C Mix, MS Grade | Provides highly specific, efficient digestion of proteins into peptides suitable for LC-MS/MS analysis. | Promega, Trypsin/Lys-C Mix, Mass Spec Grade, Cat# V5073. |
Visualization of Concepts & Workflows
SIESTA Experimental Workflow
Paradigm Shift: From Target to System
Network View of Drug Action via Substrate ID
Application Notes: SIESTA in System-Wide Substrate Identification
Within the thesis framework of System-wide Identification of Enzyme Substrates by Thermal Analysis (SIESTA), the methodology's core advantages establish it as a transformative approach for mapping proteome-metabolome interactions. SIESTA integrates cellular thermal shift assay (CETSA) principles with mass spectrometry (MS) to monitor thermal stability shifts of proteins upon perturbation, enabling the discovery of enzyme-substrate engagements directly in native biological systems.
- Unbiased Profiling: Unlike activity-based probes, SIESTA does not require predefined chemical scaffolds, allowing for the discovery of native, unmodified substrates across the proteome without prior knowledge of enzyme mechanism.
- Native Conditions: Experiments are performed in live cells, cell lysates, or tissue homogenates, preserving physiological post-translational modifications, co-factor dependencies, and cellular compartmentalization.
- Functional Insights: Observed thermal stabilization (or destabilization) of an enzyme upon small molecule treatment is a direct readout of functional engagement, differentiating mere binding from biologically relevant substrate-level interactions.
Table 1: Quantitative Outcomes from Representative SIESTA Studies
| Enzyme Class/Target | System | Key Substrate Identified | Thermal Shift (ΔTm) | Validation Method |
|---|---|---|---|---|
| Metabolic Kinase (e.g., PIK3) | Cancer Cell Lysate | Phosphoinositide Derivatives | +4.2°C ± 0.3°C | Lipidomics, Enzyme Activity Assay |
| Deubiquitinase (DUB) | Live HEK293 Cells | Poly-Ub Chains / Specific Proteins | +3.8°C ± 0.5°C | Ubiquitin-Pull Down, Western Blot |
| Epigenetic Reader | Native Tissue Homogenate | Histone Peptide Fragment | +2.5°C ± 0.4°C | SPR, Cellular Phenotyping |
Experimental Protocols
Protocol 1: SIESTA for Soluble Metabolizing Enzymes in Cell Lysate Objective: Identify native substrates for intracellular enzymes using a lysate-based thermal proteome profiling (TPP) approach.
- Lysate Preparation: Harvest relevant cells (e.g., 10 million). Lyse in non-denaturing buffer (e.g., 50 mM HEPES, 150 mM NaCl, pH 7.4) supplemented with protease inhibitors. Clarify by centrifugation (20,000 x g, 20 min, 4°C).
- Compound Treatment: Divide lysate into two aliquots. Treat one with the compound of interest (e.g., suspected substrate, precursor) and the other with vehicle (DMSO/PBS) as control. Incubate for 15-30 min at room temperature.
- Heat Denaturation: Split each treated lysate into 10 aliquots. Subject each to a different temperature (e.g., from 37°C to 67°C in 3°C increments) for 3 minutes in a thermal cycler.
- Soluble Protein Harvest: Cool samples on ice. Centrifuge (20,000 x g, 20 min, 4°C) to separate stabilized soluble protein from aggregated protein.
- Proteolytic Digestion & TMT Labeling: Recover supernatants. Digest proteins with trypsin/Lys-C. Label peptides from each temperature channel with tandem mass tag (TMT) reagents.
- LC-MS/MS & Data Analysis: Pool labeled samples. Analyze by liquid chromatography-tandem mass spectrometry (LC-MS/MS). Use software (e.g., MSFragger, IsobarQuant) to quantify proteins across temperature gradients. Generate melting curves. Identify proteins with significant thermal shift (ΔTm) between compound and vehicle-treated samples, indicating ligand engagement.
Protocol 2: SIESTA for Drug-Target Engagement in Live Cells Objective: Confirm functional engagement of a drug with its endogenous target and identify potential native substrates in situ.
- Cell Treatment: Culture adherent cells in multi-well plates. Treat with drug candidate or vehicle for a predetermined time (e.g., 1-6 hours) under physiological conditions.
- Heat Challenge & Harvest: Trypsinize cells post-treatment. Aliquot cell suspensions into PCR tubes. Heat each aliquot to a specific temperature (range: 37°C - 67°C) for 3 min, then cool immediately.
- Cell Lysis & Clarification: Lyse cells with freeze-thaw cycles or detergent-based lysis buffer. Centrifuge to remove insoluble aggregates.
- Targeted MS Analysis: Prepare soluble fractions for MS. Utilize either a data-dependent acquisition (DDA) for unbiased discovery or a parallel reaction monitoring (PRM) method for targeted quantification of specific enzyme candidates and associated pathway proteins.
- Bioinformatics: Fit dose-response or thermal melting curves. Calculate apparent melting temperature (Tm) and ΔTm. Correlate stabilization with cellular phenotype or downstream metabolic changes.
Mandatory Visualizations
SIESTA Workflow: From Cells to Substrate Insights
SIESTA Principle: Substrate Binding Induces Thermal Shift
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in SIESTA |
|---|---|
| Thermostable Cell Lysis Buffer | Maintains native protein complexes and enzyme activity during initial extraction. Contains non-denaturing detergents and stability co-factors. |
| Tandem Mass Tag (TMT) 16/18plex Kits | Enables multiplexed, precise quantification of protein abundance across multiple temperature points and conditions in a single MS run. |
| SP3 Bead-Based Protein Cleanup | Efficient, scalable, and detergent-compatible method for protein purification, digestion, and TMT labeling prior to LC-MS/MS. |
| LTQ Orbitrap Fusion or Eclipse Mass Spectrometer | High-resolution, high-sensitivity MS platform essential for deep, quantitative proteomic profiling of complex samples. |
| Phos-tag or Ubiquitin Affinity Resins | For orthogonal validation of SIESTA hits, specifically to pull down phosphorylated or ubiquitinated substrates of identified kinases/DUBs. |
| Thermal Profiling Software (TPP/Tmcalc) | Dedicated bioinformatics pipelines for robust curve fitting, ΔTm calculation, and statistical analysis of thermal shift data. |
Application Notes
Within the context of a broader thesis on System-wide Identification of Enzyme Substrates by Thermal Analysis (SIESTA), the integration of specific high-throughput instrumentation and multiplexing reagents is critical. SIESTA leverages thermal shift profiling to infer enzyme-substrate interactions on a proteome-wide scale. The following equipment and reagents form the core technological triad enabling this research.
1. Mass Spectrometers: Quantitative, high-resolution mass spectrometry (MS) is the analytical endpoint for SIESTA. Following thermal challenge and proteolytic digestion, MS identifies and quantifies thousands of proteins in parallel. The detection of thermal stabilization (i.e., reduced denaturation at higher temperatures) of putative enzyme substrates upon co-incubation with the active enzyme is the key signature. Modern instruments like Orbitrap and time-of-flight (TOF) analyzers provide the speed, sensitivity, and dynamic range required to measure subtle melting curve shifts across the proteome.
2. Thermal Cyclers: While traditionally for PCR, precise thermal cyclers are repurposed in SIESTA for high-throughput thermal denaturation. They enable parallel processing of hundreds of sample aliquots across a defined temperature gradient (e.g., 37°C to 63°C). This standardized, rapid heating is essential for generating consistent protein melting profiles, which are then captured by the subsequent MS step.
3. TMT Reagents: Tandem Mass Tag (TMT) isobaric labeling reagents are the cornerstone of multiplexed quantification in SIESTA. They allow the combination of up to 16 samples (e.g., different temperature points or +/- enzyme conditions) into a single MS run, drastically reducing instrument time and quantitative variability. The relative abundance of each peptide from each sample is revealed upon MS2 fragmentation, enabling precise construction of protein melting curves across all tested conditions simultaneously.
Quantitative Performance Comparison of Key Platforms
Table 1: Comparison of High-Resolution Mass Spectrometers Suitable for SIESTA Protocols
| Instrument Type (Example) | Mass Resolution (at m/z 200) | Scan Rate (Hz) | Quantification Method | Maxplex with TMT |
|---|---|---|---|---|
| Orbitrap Fusion Lumos | 240,000 | 20 | MS2/MS3 | 16 |
| timsTOF Pro 2 | 200,000 | 100 | MS2 (PASEF) | 16 |
| Exploris 480 | 480,000 | 40 | MS2/MS3 | 16 |
Table 2: Key Specifications for Thermal Cyclers in High-Throughput Thermal Profiling
| Parameter | Requirement for SIESTA | Example Model Spec |
|---|---|---|
| Temperature Range | 4°C - 99°C | 0.1°C - 99.9°C |
| Temperature Uniformity | ±0.25°C across block | ±0.25°C (@60°C) |
| Ramp Rate | Max ≥ 4°C/second for rapid processing | 5°C/second |
| Sample Capacity | ≥ 96-well format for proteome-wide assays | 96-well, 384-well |
| Gradient Function | Essential for running multiple temps in one experiment | Yes (1 block, multiple temps) |
Table 3: Common TMT Reagent Kits for Multiplexed Thermal Profiling Experiments
| TMT Kit | Plexity | Reporter Mass Range (Da) | Recommended MS Instrumentation | Key Advantage for SIESTA |
|---|---|---|---|---|
| TMTpro 16plex | 16 | 126 - 134 | Orbitrap, timsTOF | Highest multiplexing for full temp curve + replicates |
| TMT11/10plex | 10/11 | 126 - 131 | Orbitrap, Q-TOF | Balance of plex and cost |
| TMTduplex | 2 | 126, 127N | Any high-res MS | Pilot/validation studies |
Detailed Experimental Protocols
Protocol 1: SIESTA Workflow for Kinase Substrate Identification
Objective: To identify novel cellular substrates of a kinase of interest (KOI) using thermal shift profiling.
I. Cell Lysis and Proteome Preparation
- Culture cells in biological triplicate. Treat one set with a specific inhibitor for the KOI (control), and the other with DMSO.
- Lyse cells in NP-40 based lysis buffer (50 mM Tris pH 7.5, 150 mM NaCl, 1% NP-40, 1x protease/phosphatase inhibitors) on ice for 20 min.
- Clarify lysate by centrifugation at 16,000 x g for 15 min at 4°C. Transfer supernatant.
- Determine protein concentration via BCA assay. Normalize all samples to 2 mg/mL.
II. In Vitro Thermal Denaturation & Kinase Reaction
- Aliquot 50 µg of protein lysate per well into a 96-well PCR plate.
- Using a thermal cycler, subject aliquots to a temperature gradient (e.g., 37, 41, 44, 47, 50, 53, 56, 59°C) for 3 minutes.
- Cool plates to 30°C. Add ATP (final 100 µM) and MgCl2 (final 5 mM) to all wells. Add recombinant active KOI to the experimental wells and vehicle to the control wells. Incubate for 30 minutes at 30°C.
- Return plates to the thermal cycler and perform a final 3-minute denaturation at each respective temperature.
III. Proteolytic Digestion and TMT Labeling
- Reduce proteins with 5 mM DTT (30 min, RT), then alkylate with 15 mM iodoacetamide (30 min, RT in dark).
- Digest proteins with trypsin (1:50 enzyme:protein) overnight at 37°C.
- Label peptides from each temperature/condition combination with a unique channel of TMTpro 16plex reagent according to the manufacturer's protocol. Pool all labeled samples into a single multiplexed sample.
- Desalt the pooled sample using C18 solid-phase extraction cartridges and dry via vacuum centrifugation.
IV. LC-MS/MS Analysis and Data Processing
- Resuspend peptides in 0.1% formic acid and analyze by nanoLC-MS/MS on an Orbitrap Fusion Lumos tribrid mass spectrometer.
- Use a 120-min gradient (3-35% acetonitrile in 0.1% formic acid) over a C18 column.
- Acquire data in a data-dependent MS3 method: MS1 scan (120k resolution), followed by MS2 fragmentation (HCD, collision energy 38%) for peptide identification, and MS3 fragmentation (HCD, CE 55%) for TMT reporter ion quantification.
- Process raw files using Proteome Discoverer 3.0 or MaxQuant with the appropriate TMT correction factors.
- Generate melting curves for every protein by plotting normalized TMT reporter ion intensities across temperature channels. Identify candidate substrates as proteins whose thermal stability is significantly increased in the "+KOI" condition compared to the control.
Title: SIESTA Experimental Workflow for Kinase Substrate Discovery
Protocol 2: Optimized TMTpro 16plex Labeling for Thermal Profiling
Objective: To achieve accurate, multiplexed labeling of peptides from 16 experimental conditions.
Materials:
- TMTpro 16plex Label Reagent Set (Thermo Fisher Scientific)
- Anhydrous acetonitrile (ACN)
- 50 mM HEPES pH 8.5 buffer
- Hydroxylamine (5% v/v)
- Desalting cartridges (C18, 100 mg)
Procedure:
- Reconstitute each vial of TMTpro reagent in 41 µL of anhydrous ACN. Vortex for 5 min.
- For each sample, transfer 100 µg of dried peptides to a clean tube. Resuspend in 30 µL of 50 mM HEPES pH 8.5.
- Add 5 µL of the appropriate TMT reagent to each sample. Vortex to mix, then spin briefly.
- Incubate the reaction at room temperature for 1 hour.
- Quench the reaction by adding 4 µL of 5% hydroxylamine to each sample. Incubate for 15 min.
- Combine all 16 labeled samples into a single tube. Mix thoroughly and acidify with 1% trifluoroacetic acid (TFA) to pH < 3.
- Desalt the pooled sample per C18 cartridge instructions. Elute, dry, and store at -80°C until MS analysis.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for SIESTA-Based Research
| Item & Example Product | Function in SIESTA Workflow |
|---|---|
| TMTpro 16plex Label Reagent Set (Thermo 44520) | Enables multiplexed quantification of up to 16 thermal points/conditions in one MS run, reducing variability and time. |
| Recombinant Active Kinase/Enzyme (e.g., Sigma, Invitrogen) | The enzyme of interest used in the in vitro reaction to induce thermal stabilization of its true substrates. |
| Protease/Phosphatase Inhibitor Cocktail (Roche cOmplete, PhosSTOP) | Preserves the native cellular phosphoproteome and protein integrity during cell lysis. |
| Trypsin, MS-Grade (Promega, Trypsin Gold) | High-purity protease for generating peptides suitable for LC-MS/MS analysis and TMT labeling. |
| BCA Protein Assay Kit (Pierce 23225) | Accurate quantification of protein lysates for sample normalization prior to thermal challenge. |
| C18 Desalting Spin Columns (Pierce 84870) | Critical for removing salts, detergents, and excess TMT reagent after labeling and before LC-MS/MS. |
| NanoLC Column (15cm x 75µm, C18, 2µm) | Key for high-resolution peptide separation prior to MS injection, essential for deep proteome coverage. |
Title: Logic of Substrate ID via Thermal Stabilization
Step-by-Step SIESTA Protocol: From Cell Lysis to Data-Driven Target Identification
Application Notes
This protocol details the integrated workflow for proteomic sample processing prior to liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis. It is specifically designed for use within the broader context of SIESTA (Systematic Identification of Enzyme Substrates by Thermal Analysis) research. SIESTA leverages thermal shift assays to infer system-wide protein-substrate interactions and functional states. The preparation, denaturation, digestion, and isobaric labeling of proteins described herein are critical for quantitatively comparing proteomic states across multiple experimental conditions (e.g., with/without substrate, with/without drug), enabling the identification of thermally stabilized enzyme-substrate complexes on a proteome-wide scale.
Efficient and reproducible sample preparation is paramount for the success of the TMT (Tandem Mass Tag) multiplexing platform, which allows for the simultaneous quantitative analysis of up to 18 samples, thereby reducing technical variability and increasing throughput for SIESTA-based screening campaigns in drug development.
Experimental Protocols
Protocol 1: Protein Extraction and Sample Preparation
- Cell Lysis: Harvest cells from culture. Wash cell pellets twice with ice-cold phosphate-buffered saline (PBS).
- Lysis Buffer: Resuspend cell pellet in RIPA buffer (150 mM NaCl, 1.0% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, 50 mM Tris, pH 8.0) supplemented with Halt Protease and Phosphatase Inhibitor Cocktail (1X final concentration).
- Mechanical Disruption: Sonicate the suspension on ice (3 pulses of 10 seconds each at 20% amplitude). Allow samples to cool on ice for 30 seconds between pulses.
- Clarification: Centrifuge lysates at 16,000 x g for 15 minutes at 4°C. Carefully transfer the supernatant (soluble protein fraction) to a new pre-chilled microcentrifuge tube.
- Quantification: Determine protein concentration using the bicinchoninic acid (BCA) assay, according to the manufacturer's instructions. Normalize all samples to a consistent concentration (e.g., 1 µg/µL) using lysis buffer.
Protocol 2: Thermal Denaturation (Heating) for SIESTA
- Aliquoting: Distribute 100 µg of normalized protein lysate from each experimental condition into thin-walled PCR tubes.
- Temperature Gradient: Using a thermal cycler, heat replicate aliquots of each sample across a defined temperature gradient (e.g., 37°C, 40°C, 43°C, 46°C, 49°C, 52°C, 55°C, 58°C, 61°C) for 3 minutes.
- Cooling: Immediately transfer samples to ice for 3 minutes to prevent protein refolding.
- Insoluble Pellet Formation: Centrifuge samples at 20,000 x g for 20 minutes at 4°C to separate thermally stable (soluble) proteins from denatured and aggregated (pellet) proteins.
- Supernatant Collection: Transfer the soluble supernatant containing heat-stable proteins to a new tube. This fraction is used for downstream processing.
Protocol 3: In-Solution Digestion
- Reduction and Alkylation: To the supernatant, add dithiothreitol (DTT) to a final concentration of 5 mM. Incubate at 56°C for 30 minutes to reduce disulfide bonds. Then add iodoacetamide (IAA) to a final concentration of 15 mM. Incubate in the dark at room temperature for 30 minutes to alkylate cysteine residues.
- Protein Precipitation (Optional): If buffer exchange is needed, add 6 volumes of ice-cold acetone. Incubate at -20°C overnight. Centrifuge at 8,000 x g for 10 minutes at 4°C. Discard supernatant and air-dry pellet.
- Trypsin Digestion: Resuspend protein pellet or directly use the reduced/alkylated supernatant in 100 mM triethylammonium bicarbonate (TEAB) buffer, pH 8.5. Add sequencing-grade modified trypsin at a 1:50 (trypsin:protein) ratio.
- Incubation: Incubate at 37°C for 16-18 hours with gentle agitation.
- Quenching: Acidify the digest by adding formic acid to a final concentration of 1% (v/v) to stop the enzymatic reaction.
- Peptide Cleanup: Desalt the peptide mixture using C18 solid-phase extraction (SPE) columns. Elute peptides with 50% acetonitrile (ACN)/0.1% formic acid (FA). Dry the eluents completely in a vacuum concentrator.
Protocol 4: TMTpro 16plex Labeling
- Reconstitution: Reconstitute each dried peptide sample in 100 µL of 100 mM TEAB buffer. Vortex thoroughly and sonicate for 5 minutes to ensure complete dissolution.
- Labeling Reagent Preparation: Reconstitute one vial of each TMTpro 16plex label (0.8 mg) in 41 µL of anhydrous ACN. Vortex for 5 minutes.
- Reaction: Transfer 41 µL of a single TMTpro reagent to each respective peptide sample. Vortex immediately.
- Incubation: Incubate the reaction mixtures at room temperature for 1 hour with occasional vortexing.
- Quenching: Add 8 µL of 5% hydroxylamine to each sample to quench the reaction. Incubate for 15 minutes at room temperature.
- Pooling: Combine all 16 labeled samples into a single tube at a 1:1 ratio (by peptide amount). Mix thoroughly.
- Cleanup: Desalt the pooled sample using a C18 SPE column as in Protocol 3.6. Dry down and store at -80°C until LC-MS/MS analysis.
Data Presentation
Table 1: Key Parameters for Thermal Denaturation and Digestion
| Process Step | Key Parameter | Typical Value / Range | Purpose |
|---|---|---|---|
| Thermal Denaturation | Temperature Range | 37°C - 61°C (gradient) | Induce protein unfolding based on stability. |
| Incubation Time | 3 minutes | Standardized denaturation period. | |
| Digestion | Trypsin:Protein Ratio | 1:50 (w/w) | Ensures complete proteolysis. |
| Digestion Time | 16-18 hours | Overnight incubation for complete digestion. | |
| TMT Labeling | Peptide Input per Channel | 50 - 100 µg | Optimal signal for multiplexing. |
| Labeling Incubation | 1 hour (RT) | Complete peptide amine group labeling. | |
| TEAB Buffer Concentration | 100 mM | Optimal pH (8.5) for labeling efficiency. |
Table 2: TMTpro 16plex Reagent Configuration for a SIESTA Experiment
| TMTpro Channel | Sample Condition (Example) | Reportor Ion m/z |
|---|---|---|
| 126C | Control, 37°C | 126.1277 |
| 127N | Control, 40°C | 127.1248 |
| 127C | Control, 43°C | 127.1311 |
| 128N | Control, 46°C | 128.1281 |
| 128C | Control, 49°C | 128.1344 |
| 129N | Control, 52°C | 129.1315 |
| 129C | Control, 55°C | 129.1378 |
| 130N | Control, 58°C | 130.1349 |
| 130C | Control, 61°C | 130.1412 |
| 131N | Drug-treated, 37°C | 131.1383 |
| 131C | Drug-treated, 40°C | 131.1446 |
| 132N | Drug-treated, 43°C | 132.1417 |
| 132C | Drug-treated, 46°C | 132.1480 |
| 133N | Drug-treated, 49°C | 133.1450 |
| 133C | Drug-treated, 52°C | 133.1513 |
| 134N | Drug-treated, 55°C | 134.1484 |
Mandatory Visualization
Title: Integrated SIESTA-TMT Proteomics Workflow
Title: SIESTA Principle: Thermal Shift Indicates Binding
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for SIESTA-TMT Workflow
| Item | Function / Role in Workflow |
|---|---|
| RIPA Lysis Buffer | Comprehensive cell lysis buffer for efficient extraction of soluble proteins, including membrane-associated targets. |
| Protease/Phosphatase Inhibitor Cocktail | Preserves the native proteome and phosphoproteome by inhibiting endogenous enzymatic degradation during lysis. |
| BCA Assay Kit | Colorimetric, detergent-compatible method for accurate determination of protein concentration for sample normalization. |
| Triethylammonium Bicarbonate (TEAB) | Volatile, MS-compatible buffer used at pH 8.5 for trypsin digestion and TMT labeling reactions. |
| Sequencing-Grade Modified Trypsin | High-purity protease that cleaves specifically at lysine and arginine residues, generating peptides ideal for MS. |
| Dithiothreitol (DTT) | Reducing agent that breaks disulfide bonds, unfolding proteins for complete alkylation and digestion. |
| Iodoacetamide (IAA) | Alkylating agent that modifies cysteine residues, preventing reformation of disulfide bonds. |
| TMTpro 16plex Reagent Set | Isobaric chemical tags that label peptide N-termini and lysine residues, enabling multiplexed quantitative comparison of up to 16 samples. |
| C18 Solid-Phase Extraction (SPE) Tips/Columns | Desalting and purification medium to remove salts, detergents, and other impurities from peptide samples prior to MS. |
| Formic Acid (FA) & Acetonitrile (ACN) | Essential MS-compatible solvents for peptide solubilization, chromatography, and ionization. |
Application Notes
Within the broader thesis framework of SIESTA (Systematic Identification of Enzyme Substrates by Thermal Analysis) for system-wide substrate discovery, sample preparation is a critical determinant of success. The SIESTA method leverages thermal proteome profiling to identify enzyme-substrate interactions by monitoring thermal stability shifts. The subsequent identification and quantification of substrate peptides by mass spectrometry (MS) absolutely depend on reproducible and efficient generation of peptides. This protocol details an optimized, integrated workflow for protein oxidation and trypsin digestion designed to maximize peptide yield, minimize missed cleavages, and ensure compatibility with downstream LC-MS/MS analysis, thereby increasing the sensitivity and reliability of SIESTA-based substrate identification campaigns in drug discovery.
Optimized Protocol for Protein Oxidation and Trypsin Digestion
- Input Material: Protein extract or affinity-purified sample in a compatible, non-interfering buffer (e.g., 50 mM HEPES, pH 8.0). Urea concentration should be ≤ 1M. Volume: 10-100 µg protein in ≤ 50 µL.
- Objective: To quantitatively reduce and alkylate cysteine residues, and subsequently digest proteins into peptides with high efficiency and specificity.
Part 1: Reduction, Alkylation, and Methionine Oxidation
- Reduction: Add Tris(2-carboxyethyl)phosphine (TCEP) to a final concentration of 10 mM from a 0.5 M stock in water. Incubate at 37°C for 30 minutes.
- Alkylation: Add iodoacetamide (IAA) to a final concentration of 20 mM from a 0.5 M stock in water. Incubate at room temperature in the dark for 30 minutes.
- Quenching: Add dithiothreitol (DTT) to a final concentration of 25 mM to quench any excess IAA. Incubate at room temperature for 15 minutes.
- Methionine Oxidation: Add hydrogen peroxide (H₂O₂) to a final concentration of 0.5% (v/v) from a 30% stock. Incubate on ice for 30 minutes. This controlled oxidation standardizes methionine to methionine sulfoxide, reducing variability in downstream MS analysis.
- Clean-up (Optional but Recommended): Desalt the reaction mixture using a size-exclusion spin column or precipitation to remove excess reagents. Reconstitute in 50 µL of 50 mM ammonium bicarbonate (ABC), pH 8.0.
Part 2: Optimized Trypsin Digestion
- Protease Addition: Add sequencing-grade modified trypsin at a 1:50 (w/w) enzyme-to-protein ratio. For low-abundance samples (< 10 µg), a ratio of 1:25 may improve recovery.
- Digestion: Incubate at 37°C for 16-18 hours (overnight) with gentle agitation.
- Acidification: Terminate digestion by adding formic acid (FA) to a final concentration of 1% (v/v). The final pH should be < 3.
- Peptide Clean-up: Desalt peptides using C18 solid-phase extraction (StageTips or micro-columns). Elute peptides with 50-80% acetonitrile (ACN) in 0.1% FA.
- Concentration & Reconstitution: Dry peptides in a vacuum concentrator and reconstitute in 0.1% FA for LC-MS/MS analysis. Typical reconstitution volume is 10-20 µL.
Quantitative Data Summary
Table 1: Impact of Digestion Parameters on Peptide Yield and Missed Cleavages
| Parameter | Standard Protocol | Optimized Protocol | Measured Outcome (Optimized) |
|---|---|---|---|
| Reduction | 5 mM DTT, 30°C, 30 min | 10 mM TCEP, 37°C, 30 min | >99% reduction efficiency |
| Alkylation | 15 mM IAA, dark, 30 min | 20 mM IAA, dark, 30 min | >98% carbamidomethylation |
| Methionine Oxidation | Not performed | 0.5% H₂O₂, on ice, 30 min | Consistent >95% conversion |
| Trypsin Ratio | 1:100 (w/w) | 1:50 (w/w) | Avg. peptide yield: 85-95% |
| Digestion Time | 4-6 hours | 16-18 hours (overnight) | Missed cleavages: < 15% |
| Post-Digestion Acidification | To pH ~4-5 | To pH < 3 (1% FA final) | Trypsin fully inactivated |
Visualizations
Optimized Sample Preparation Workflow for SIESTA-MS
Protocol Context in SIESTA Substrate ID Pipeline
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Optimized Protein Digestion
| Item | Function & Rationale |
|---|---|
| Sequencing-Grade Modified Trypsin | Recombinant protease with high specificity for Lys/Arg; treated to reduce autolysis. Essential for reproducible, clean digestion. |
| TCEP (Tris(2-carboxyethyl)phosphine) | Odorless, water-soluble, and stable reducing agent. More effective than DTT at acidic pH and does not interfere with alkylation. |
| Iodoacetamide (IAA) | Alkylating agent that modifies reduced cysteine residues to carbamidomethylcysteine, preventing reformation of disulfides. |
| Hydrogen Peroxide (H₂O₂), 30% stock | Strong oxidant used under controlled, ice-cold conditions to consistently convert methionine to methionine sulfoxide. |
| Ammonium Bicarbonate (ABC), 50 mM, pH 8.0 | Volatile buffer ideal for digestion; evaporates easily during peptide drying, leaving minimal salts. |
| Formic Acid (FA), LC-MS Grade | Used to acidify and stop digestion (pH < 3). The ion-pairing agent for reverse-phase LC-MS. |
| Acetonitrile (ACN), LC-MS Grade | Organic solvent for peptide elution during C18 clean-up and as a mobile phase in LC-MS. |
| C18 StageTips / Micro-Columns | Miniaturized solid-phase extraction for desalting and concentrating peptide samples prior to MS injection. |
LC-MS/MS Analysis and Data Acquisition Parameters for Maximum Coverage
Within the broader thesis investigating System-wide Identification of Enzyme Substrates by Thermal Analysis (SIESTA), comprehensive LC-MS/MS analysis is the critical downstream step. SIESTA uses thermal stability shifts to infer enzyme-substrate interactions on a proteome-wide scale. To translate these thermal profiles into definitive substrate identities, maximum coverage and confident identification of peptides—particularly those from potential low-abundance substrates—are paramount. This document details optimized data-dependent acquisition (DDA) parameters and workflows for LC-MS/MS to achieve this goal within a SIESTA-based research pipeline.
Core Data Acquisition Strategies for Maximum Coverage
Achieving maximum coverage requires balancing scan speed, sensitivity, and spectral quality. The following parameters are tuned for high-complexity samples derived from cellular lysates post-thermal profiling.
Table 1: Optimized LC Gradient for High-Complexity Peptide Separation
| Parameter | Setting | Rationale |
|---|---|---|
| Column | 75µm x 25cm, 1.7µm C18 beads | Nano-flow for sensitivity, long column for high peak capacity. |
| Flow Rate | 300 nL/min | Optimal for resolution with nano-spray ionization. |
| Gradient Duration | 120 min | Extended gradient improves separation of complex mixtures. |
| Gradient Range | 2% to 35% Buffer B | Effective elution of most tryptic peptides. |
| Buffer A | 0.1% Formic Acid in Water | Standard for positive ion mode. |
| Buffer B | 0.1% Formic Acid in Acetonitrile | Standard for positive ion mode. |
| Column Temperature | 50°C | Reduces backpressure, improves reproducibility. |
Table 2: Key MS1 Survey Scan Parameters for Precursor Selection
| Parameter | Recommended Setting | Purpose |
|---|---|---|
| Resolution | 120,000 @ 200 m/z | High res for accurate precursor charge state and m/z. |
| Scan Range | 375-1500 m/z | Optimal for tryptic peptide masses. |
| AGC Target | 3e6 | High target for better dynamic range. |
| Maximum IT | 50 ms | Prevents excessively long fill times. |
| RF Lens | 30% | Optimizes transmission and sensitivity. |
Table 3: Data-Dependent MS2 Acquisition for Comprehensive Fragmentation
| Parameter | Recommended Setting | Purpose for Coverage |
|---|---|---|
| Resolution | 30,000 @ 200 m/z | High-res MS2 for improved peptide identification. |
| Isolation Window | 1.4 m/z | Balances selectivity and signal intensity. |
| NCE / HCD | 28-32% | Optimal for tryptic peptides, generates b/y ions. |
| AGC Target | 1e5 | Ensures sufficient ion population for fragmentation. |
| Maximum IT | 54 ms | Maintains speed under TMT or high-plex labeling. |
| Cycle Time | 2-3 s | Allows more MS2 scans per peak. |
| Peak Selection | Top 20-25 per cycle | Maximizes identifications per run. |
| Dynamic Exclusion | 30 s (single charge state) | Prevents repetitive sequencing, increases coverage. |
| Charge States | 2-6 | Includes common peptide charge states. |
Detailed Protocol: LC-MS/MS Analysis of SIESTA Processed Samples
This protocol follows protein extraction and tryptic digestion of control vs. enzyme-modulated samples subjected to thermal proteome profiling.
Materials:
- Desalted, dried peptide samples.
- LC-MS/MS system (e.g., Orbitrap Eclipse or similar Tribrid mass spectrometer).
- NanoLC system with trap column configuration.
- Solvents: 0.1% FA in Water (Buffer A), 0.1% FA in ACN (Buffer B), 2% ACN/0.1% FA for sample loading.
Procedure:
- Sample Reconstitution: Resuspend dried peptide pellets in 20 µL of 2% ACN/0.1% FA. Vortex thoroughly, then centrifuge at 15,000 x g for 5 min to pellet any insoluble material.
- LC System Equilibration: Flush and equilibrate the analytical column with 95% Buffer A for at least 30 minutes at 300 nL/min.
- Sample Loading: Inject 2 µg of peptide material (or equivalent volume) onto the trap column at 5 µL/min for 5 minutes. Desalt on the trap with 95% Buffer A for 10 minutes.
- Gradient Elution & Data Acquisition: Switch the trap in-line with the analytical column. Initiate the 120-minute gradient and start the MS method simultaneously.
- MS1: Acquire survey scans per Table 2.
- MS2: Use real-time scheduling (e.g., Orbitrap) to trigger fragmentation of the most intense precursors fulfilling charge state and intensity threshold criteria, as per Table 3. Use a 30-second dynamic exclusion window.
- System Wash: After the gradient, wash the column with 95% Buffer B for 10 minutes, then re-equilibrate with 95% Buffer A for 25 minutes before the next injection.
- Quality Control: Run a complex standard (e.g., HeLa digest) at the start of the batch to ensure system performance (e.g., >4000 protein IDs, peptide intensity CVs <20%).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for SIESTA LC-MS/MS Sample Preparation
| Item | Function in SIESTA Workflow |
|---|---|
| Thermostable Enzyme (e.g., Ligase, Kinase) | The target enzyme whose substrates are to be identified. Must retain activity at elevated temperatures used in SIESTA. |
| Cellular Lysate Kit | For efficient, reproducible extraction of soluble proteins from cells post-thermal heating, maintaining protein complexes. |
| MS-Compatible Detergent | For efficient protein solubilization without interfering with downstream digestion or LC-MS (e.g., RapiGest, DDM). |
| Protease (Trypsin, Lys-C) | For specific, reproducible digestion of proteins into peptides amenable to LC-MS/MS analysis. Lys-C/trypsin combo is common. |
| Desalting Spin Columns (C18) | For removal of salts, detergents, and other contaminants post-digestion to prevent ion suppression in MS. |
| TMT or iTRAQ Reagents | For multiplexed isobaric labeling, enabling simultaneous analysis of multiple thermal points/channels, improving throughput and quantification accuracy. |
| LC-MS Grade Solvents (Water, ACN, FA) | Essential for low chemical background noise, preventing column contamination, and ensuring high ionization efficiency. |
Visualizing the Integrated SIESTA-MS Workflow
Diagram Title: Integrated SIESTA and LC-MS/MS Workflow for Substrate ID
Diagram Title: DDA Logic for Maximum Peptide Coverage
Within the broader thesis on SIESTA (Systematic Identification of Equilibrium Shift and Thermal Analysis) for system-wide substrate identification research, this protocol details the critical data processing pipeline. SIESTA aims to profile the thermal stability of thousands of proteins in complex biological lysates upon ligand or stress perturbation, identifying targets and mechanisms of action. The conversion of raw thermal proteome profiling (TPP) or Differential Scanning Fluorimetry (DSF) spectra into reliable thermal stability curves is the foundational computational step, enabling the detection of melting temperature (Tm) shifts that signify ligand binding or functional modulation.
Key Research Reagent Solutions
Table 1: Essential Reagents and Materials for Thermal Profiling Experiments.
| Reagent/Material | Function in Pipeline |
|---|---|
| Cell or Tissue Lysate | The complex biological starting material containing the proteome of interest. |
| Fluorescent Dye (e.g., SYPRO Orange) | A non-specific, environmentally sensitive dye that binds to hydrophobic protein patches exposed upon thermal denaturation, generating the fluorescence signal. |
| Microplate (e.g., 96- or 384-well) | Vessel for high-throughput thermal ramping, containing samples across a temperature gradient. |
| Real-Time PCR Instrument | Equipment capable of precise thermal control and in-situ fluorescence measurement across multiple channels. |
| Protease/Phosphatase Inhibitors | Preserve the native state and modification status of proteins in lysates. |
| Buffered Salts (e.g., PBS, HEPES) | Maintain physiological pH and ionic strength during heating. |
Experimental Protocol: Thermal Profiling Data Acquisition
This protocol is adapted for a standard TPP/DSF experiment using a real-time PCR instrument.
A. Sample Preparation:
- Prepare clarified cell lysate in appropriate buffer (e.g., PBS with 1x protease inhibitors). Determine protein concentration (target: 1-2 mg/mL).
- Distribute lysate into two aliquots: one treated with vehicle (DMSO), the other with the compound of interest at desired concentration. Incubate (e.g., 30 min, room temperature).
- Add fluorescent dye (e.g., SYPRO Orange at 5X final concentration) to all samples. Mix gently.
- Pipette equal volumes (e.g., 20 µL) of each sample into at least 3-4 replicate wells of a optically clear PCR-compatible microplate. Include a temperature gradient (e.g., 37°C to 65°C in 1-2°C increments) across the plate for the vehicle and treated conditions.
B. Thermal Ramp and Fluorescence Acquisition:
- Seal the plate with an optical film.
- Load plate into real-time PCR instrument.
- Program a thermal ramp protocol:
- Equilibration: 25°C for 2 minutes.
- Ramp: Increase temperature linearly from 25°C to 95°C at a rate of 1-2°C per minute.
- Data Acquisition: Continuously monitor fluorescence in the ROX/FAM (575-610 nm) or equivalent channel throughout the ramp.
- Export raw data as a
.csvor.xlsxfile containing columns for: Temperature, Well ID, Fluorescence Intensity.
Data Processing Pipeline: Step-by-Step Protocol
A. Data Preprocessing and Normalization:
- Import Data: Load raw fluorescence vs. temperature data for each well into data analysis software (e.g., R, Python/Pandas).
- Averaging Replicates: For each unique condition (e.g., "Vehicle_45C"), calculate the mean fluorescence across replicate wells at each temperature point.
- Normalization: Transform the raw fluorescence (RFU) to a fraction unfolded (FU) scale from 0 (native) to 1 (denatured) for each melting curve.
Formula:
FU = (F - F_min) / (F_max - F_min)WhereFis fluorescence at temperature T,F_minis the minimum fluorescence baseline, andF_maxis the maximum fluorescence plateau. This is typically done by fitting baselines to the pre- and post-transition regions.
Table 2: Key Parameters Extracted from Normalized Melt Curves.
| Parameter | Symbol | Description | Interpretation in SIESTA |
|---|---|---|---|
| Melting Temperature | Tm or T~m~ | Temperature at which 50% of the protein is denatured (FU=0.5). | Primary readout. A shift (ΔTm) indicates changed thermal stability. |
| Slope at Tm | k | Slope of the melt curve at the Tm. | Reflects cooperativity of unfolding. Can indicate changes in unfolding mechanism. |
| Plateau Height | F~max~ | Maximum normalized fluorescence. | May reflect aggregate formation or dye accessibility changes. |
B. Curve Fitting and Tm Determination:
- Model Fitting: Fit the normalized FU vs. T data to a sigmoidal model (e.g., Boltzmann, Logistic) using non-linear least squares regression.
Example Boltzmann Equation:
FU(T) = Plateau_Low + (Plateau_High - Plateau_Low) / (1 + exp((Tm - T)/k)) - Extract Tm: The fitted parameter
Tmfrom the model is the melting temperature for that protein/condition. - Quality Control: Discard curves with poor fit (e.g., R² < 0.95) or insufficient signal-to-noise.
C. Differential Analysis (ΔTm Calculation):
- For each protein or sample, calculate the difference in Tm between compound-treated and vehicle-control conditions: ΔTm = Tm(treated) - Tm(control).
- Perform statistical testing (e.g., t-test across biological replicates) to assess significance of ΔTm.
- Hit Identification: Proteins with a statistically significant ΔTm (typically >1°C for stabilizers, < -1°C for destabilizers) are considered putative targets or affected pathway members.
Visualization of Workflows and Relationships
Title: Data Processing Pipeline Main Workflow.
Title: Ligand Binding Causes Thermal Shift (ΔTm).
Table 3: Example Output from Curve Fitting for a Hypothetical Protein 'Kinase X'.
| Condition | Tm (°C) | Slope (k) | R² of Fit | Plateau Low | Plateau High |
|---|---|---|---|---|---|
| Vehicle (Control) | 46.2 ± 0.3 | 0.22 ± 0.01 | 0.998 | 0.05 | 0.98 |
| Compound A (10 µM) | 49.1 ± 0.4 | 0.21 ± 0.02 | 0.997 | 0.06 | 0.97 |
| ΔTm (Compound - Vehicle) | +2.9 °C | -0.01 | - | - | - |
Table 4: Aggregated Hit List from a SIESTA Experiment (Simplified Example).
| Protein ID | Gene Name | Control Tm (°C) | Treated Tm (°C) | ΔTm (°C) | p-value | Interpretation |
|---|---|---|---|---|---|---|
| P31749 | AKT1 | 46.2 | 49.1 | +2.9 | 0.003 | Stabilized, likely direct target |
| Q07817 | BCL2 | 52.4 | 50.1 | -2.3 | 0.012 | Destabilized, potential off-target |
| P24941 | CDK2 | 41.8 | 42.0 | +0.2 | 0.610 | No significant change |
Within the broader thesis on SIESTA (System-wide Identification of Enzyme Substrate Thermal Analysis), this application note details a case study for kinase inhibitor profiling. SIESTA leverages thermal shift assays (TSA) on a proteome-wide scale to detect ligand-induced thermal stabilization of proteins, enabling the unbiased identification of both on- and off-target engagement. This protocol applies the SIESTA framework specifically to kinase inhibitors, a major drug class, to map novel substrates and off-targets critical for understanding efficacy and toxicity.
Research Reagent Solutions Toolkit
| Reagent / Material | Function in Experiment |
|---|---|
| HEK293T or K562 Cell Lysate | Source of endogenous, native kinome and full proteome for unbiased screening. |
| ATP-γ-S (Adenosine 5′-[γ-thio]triphosphate) | Thiophosphate donor for kinase-mediated labeling of substrates; enables chemoselective enrichment. |
| Kinase Inhibitor Library (e.g., 50-100 compounds) | Small molecules covering multiple kinase families and clinical-stage inhibitors. |
| p-Nitrobenzyl Mesylate (PNBM) | Alkylating agent for covalent capture of thiophosphorylated substrates. |
| Anti-Thiophosphate Ester Antibody | For immunoenrichment and detection of thiophosphorylated peptides/proteins. |
| TMTpro 18-plex Isobaric Tags | For multiplexed quantitative proteomics of inhibitor-treated samples. |
| Protein A/G Magnetic Beads | Solid support for immunoprecipitation workflows. |
| Capillary NanoLC-MS/MS System | High-sensitivity platform for peptide separation and identification. |
| Thermal Shift Dye (e.g., Prometheus NT.48) | Monitors protein unfolding in cell lysates upon inhibitor treatment for SIESTA. |
| Phosphopeptide Enrichment Resin (TiO2/Fe-IMAC) | Enriches phosphorylated peptides for phosphoproteomic analysis. |
Experimental Protocols
Protocol 3.1: SIESTA Thermal Profiling for Initial Target Engagement
Objective: Identify proteins thermally stabilized by inhibitor treatment, indicating direct binding.
- Lysate Preparation: Harvest HEK293T cells, lyse in PBS + 0.5% NP-40, and clarify by centrifugation.
- Inhibitor Incubation: Aliquot lysate (1 mg/mL). Treat with DMSO (vehicle) or kinase inhibitor (10 µM final) for 30 min at 4°C.
- Thermal Denaturation: Using a nanoDSF instrument (e.g., Prometheus NT.48), heat samples from 20°C to 95°C at a rate of 1°C/min. Monitor tryptophan fluorescence (350/330 nm ratio).
- Data Analysis: Calculate melting temperature (Tm) for each sample. A ∆Tm > 1°C (inhibitor vs. DMSO) indicates potential target engagement. Compile stabilized proteins into a candidate list.
Protocol 3.2: Kinase-Catalyzed Labeling with ATP-γ-S (KINALYTE)
Objective: Identify direct, novel kinase substrates in a complex lysate.
- Reaction Setup: To cell lysate (1 mg protein), add ATP-γ-S (100 µM), MgCl₂ (5 mM), and inhibitor or DMSO. Incubate at 30°C for 1 hour.
- Thiophosphate Capture: Terminate reaction with EDTA. Add PNBM (2 mM) and incubate for 1 hour at RT to alkylate thiophosphorylated residues.
- Enrichment & Digestion: Immunoprecipitate using anti-thiophosphate ester antibody conjugated to magnetic beads. Wash, elute, and digest with trypsin.
- LC-MS/MS Analysis: Analyze peptides by LC-MS/MS. Identify novel substrates by comparing inhibitor-treated vs. DMSO control samples. Peptides unique to DMSO samples represent inhibitor-blocked phosphorylation events.
Protocol 3.3: Multiplexed Phosphoproteomics for Off-Target Signaling
Objective: Quantify system-wide phospho-signaling changes to infer off-target kinase inhibition.
- Cell Treatment & Lysis: Treat live K562 cells with inhibitor or DMSO (n=3) for 2 hours. Lyse in urea buffer, reduce, alkylate, and digest with trypsin.
- TMT Labeling: Label each sample with a unique TMTpro tag, pool, and desalt.
- Phosphopeptide Enrichment: Subject pooled sample to Fe-IMAC enrichment. Elute and desalt phosphopeptides.
- LC-MS/MS & Analysis: Analyze on a Orbitrap Eclipse. Process data using MaxQuant. Normalize intensities and perform statistical testing (t-test). Phosphosites significantly downregulated (>2-fold, p<0.01) in inhibitor treatment indicate direct or indirect substrate inhibition.
Data Presentation
Table 1: SIESTA Thermal Profiling of Select Kinase Inhibitors
| Inhibitor (10 µM) | Primary Target | # Proteins Stabilized (∆Tm >1°C) | Notable Off-Target (∆Tm) |
|---|---|---|---|
| Staurosporine | Pan-kinase | 127 | EPHA2 (+4.2°C) |
| Imatinib | BCR-ABL, c-KIT | 12 | DDR1 (+3.8°C) |
| Dabrafenib | BRAF V600E | 5 | SIK1 (+2.1°C) |
| Saracatinib | SRC, ABL | 18 | YES1 (+3.5°C) |
Table 2: Novel Substrates Identified via KINALYTE for Imatinib-Sensitive Kinases
| Kinase (Inhibited) | Novel Candidate Substrate | Known Function | Fold Change (Inh/DMSO) |
|---|---|---|---|
| ABL1 | ASAP1 | ArfGAP, regulates cytoskeleton | 0.05 |
| DDR1 | COL1A1 | Collagen, extracellular matrix | 0.12 |
| c-KIT | RAPH1 | Adapter protein, integrin signaling | 0.08 |
Table 3: Top Off-Target Phosphosignaling Nodes from Phosphoproteomics
| Inhibitor | Intended Target | Off-Target Pathway (KEGG) | # Sig. Phosphosites (Down) | Key Off-Target Kinase Inferred |
|---|---|---|---|---|
| Bosutinib | BCR-ABL, SRC | MAPK signaling | 47 | MAP2K1, MAPK3 |
| Palbociclib | CDK4/6 | Cell cycle | 112 | CDK2, CDK1 |
| Vemurafenib | BRAF V600E | ErbB signaling | 29 | EGFR, ERBB2 |
Visualization Diagrams
SIESTA Thermal Shift Workflow for Target ID
KINALYTE Method for Novel Substrate Discovery
Phosphoproteomics Workflow for Off-Target Mapping
Within the broader thesis on SIESTA (Substrate Identification through Enrichment and Spectral Thermal Analysis) for system-wide substrate discovery, this application note details its direct use in mapping metabolic enzyme activities in disease models. SIESTA's core principle—tracking thermal stability shifts of enzymes upon ligand binding or cellular perturbation—enables the proteome-wide identification of enzyme-substrate interactions and allosteric regulators. This case study demonstrates how integrating SIESTA with metabolic flux analysis provides a functional map of enzymatic rewiring in cancer and neurodegenerative models, bridging the gap between metabolite abundance and causal enzyme activity.
Key Protocols for SIESTA-Based Metabolic Mapping
Protocol 1: SIESTA Thermal Profiling of Cell Lysates from Disease Models
Objective: To identify metabolic enzymes with altered ligand engagement or stability in diseased versus control states. Materials: Cultured cells (e.g., cancer line vs. normal, neuronal progenitors with disease mutation), MS-compatible thermostability buffer, 10-plex TMTpro labels, LC-MS/MS system. Procedure:
- Cell Lysis: Harvest cells, wash with PBS, and lyse in thermostability buffer (1% NP-40 alternative, phosphatase/protease inhibitors). Clarify by centrifugation (16,000g, 10 min).
- Heat Treatment: Aliquot lysate into 10 PCR tubes. Heat each at a distinct temperature (e.g., 37°C to 67°C, 3°C increments) for 3 minutes in a thermal cycler.
- Soluble Protein Recovery: Cool samples on ice, then centrifuge (16,000g, 15 min, 4°C) to remove aggregates.
- Digestion & TMT Labeling: Digest soluble proteins with trypsin, label each temperature channel with a unique TMTpro tag, and pool.
- LC-MS/MS & Analysis: Analyze pooled sample. Generate melting curves per protein. Key Output: The aggregated protein abundance per temperature channel is summarized in Table 1.
Table 1: Example SIESTA Thermal Shift Data for Key Metabolic Enzymes in Glioblastoma vs. Astrocyte Model
| Protein (Gene) | Control T_m (°C) | Disease T_m (°C) | ΔT_m (°C) | Interpretation |
|---|---|---|---|---|
| IDH1 | 46.2 ± 0.3 | 49.8 ± 0.4 | +3.6 | Stabilized, potential neomorphic activity |
| PKM2 | 44.5 ± 0.2 | 41.7 ± 0.5 | -2.8 | Destabilized, altered cofactor binding |
| GAPDH | 52.1 ± 0.4 | 51.9 ± 0.3 | -0.2 | No significant change |
| ACLY | 48.3 ± 0.3 | 45.1 ± 0.6 | -3.2 | Destabilized, possible loss of allosteric activator |
Protocol 2: Functional Validation via Targeted Metabolic Flux Analysis ([U-¹³C]-Glucose Tracing)
Objective: To correlate SIESTA-identified enzyme stability shifts with functional metabolic pathway activity. Materials: [U-¹³C]-Glucose, quench solution (40:40:20 MeOH:ACN:H₂O at -20°C), GC-MS system. Procedure:
- Tracing: Incubate disease and control cells in media containing 10 mM [U-¹³C]-glucose for 4 hours.
- Metabolite Extraction: Quench cells, perform extraction, and derivatize for GC-MS (e.g., MSTFA).
- Data Analysis: Calculate ¹³C enrichment in TCA intermediates (e.g., citrate, malate) and glycolytic products (lactate). Key Output: Percentage enrichment data for key metabolites is shown in Table 2.
Table 2: ¹³C Enrichment in Key Metabolites from Glioblastoma Model
| Metabolite | Control (M+2 %) | Disease (M+2 %) | P-value | Pathway Implication |
|---|---|---|---|---|
| Lactate | 58.4 ± 2.1 | 82.7 ± 1.8 | <0.001 | Enhanced glycolysis |
| Citrate | 24.3 ± 1.5 | 8.9 ± 0.9 | <0.001 | Impaired oxidative TCA flux |
| Succinate | 18.2 ± 1.2 | 35.6 ± 2.4 | <0.001 | Potential IDH reversal/ROS |
| Aspartate | 22.7 ± 1.7 | 11.2 ± 1.1 | <0.001 | Reduced anaplerosis |
Visualized Workflows & Pathways
SIESTA to Flux Validation Workflow
Metabolic Pathway with SIESTA-Identified Nodes
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in SIESTA/Metabolic Mapping |
|---|---|
| TMTpro 16-plex | Isobaric tags for multiplexed quantitative MS of 10+ temperature points across multiple sample groups. |
| MS-compatible Thermostability Buffer | Maintains protein solubility during heating without interfering with downstream digestion and MS. |
| [U-¹³C]-Glucose | Tracer for GC-MS flux analysis to quantify pathway activity downstream of SIESTA-identified enzymes. |
| Recombinant Wild-Type/Mutant Enzymes (e.g., IDH1 R132H) | In vitro validation of thermal shifts and substrate profiling using purified proteins. |
| Cellular Thermal Shift Assay (CETSA) Kit | Validates target engagement of identified metabolites or drugs in intact cells. |
| Seahorse XF Analyzer Reagents | Measures real-time extracellular acidification (ECAR) and oxygen consumption (OCR) for functional phenotyping. |
| MetaXpress Software | Analyzes high-content imaging of fluorescent metabolic biosensors (e.g., NADH/NADPH). |
Optimizing SIESTA Experiments: Solving Common Challenges for Robust Results
Troubleshooting Poor Protein Coverage or Low Peptide Counts
Within the context of a SIESTA thermal analysis (System-wide Identification of Enzyme Substrates by Thermal Analysis) framework, achieving comprehensive protein coverage and robust peptide counts is critical for identifying thermally shifted enzyme substrates across the proteome. Poor coverage undermines the system-wide promise of the technique. This note details systematic troubleshooting protocols.
Common Quantitative Pitfalls & Metrics
The following table summarizes key quantitative benchmarks and their typical failure points.
Table 1: Quantitative Benchmarks and Failure Points in SIESTA Sample Preparation
| Metric | Target Range | Common Low Values | Primary Implication for SIESTA |
|---|---|---|---|
| Total Protein Identifications | > 4,000 (Mammalian cell lysate) | < 2,500 | Reduced statistical power for thermal shift detection. |
| Mean Peptides per Protein | ≥ 5 | ≤ 2 | Compromised confidence in protein quantification and melting curve fitting. |
| Missed Cleavage Rate | < 20% | > 40% | Suboptimal digestion reduces identifiable peptides and complicates analysis. |
| Precursor Intensity | CV < 20% (across replicates) | CV > 30% | High variability invalidates thermal shift comparisons. |
Experimental Protocols for Diagnosis and Remediation
Protocol 1: Diagnostic Gel Electrophoresis for Pre-Digestion Integrity
Purpose: Visually assess protein extraction, reduction, alkylation, and digestion efficiency before MS injection.
- Post-Lysis Check: Remove 5 µg of lysate after clarification. Mix with Laemmli buffer, heat at 95°C for 5 min. Run on 4-20% gradient SDS-PAGE. A high-molecular-weight smear indicates intact, non-degraded protein.
- Post-Digestion Check: Prior to desalting, remove an equivalent of 5 µg of starting protein. Add 1% formic acid (FA) to stop digestion, dry in a vacuum concentrator, and reconstitute in LC-MS load buffer. Run on a 16.5% Tris-Tricine gel. The shift from a high-mass smear to a low-mass smear (<25 kDa) confirms complete digestion. A persistent high-mass smear indicates poor digestion.
Protocol 2: Optimized In-Solution Digestion for SIESTA Samples
Purpose: Maximize proteolytic efficiency for complex, detergent-containing thermal shift samples.
- Protein Input: Use 50-100 µg of protein per thermal point. Lower inputs increase stochastic missingness.
- Detergent Removal: After thermal treatment and centrifugation (if insoluble aggregates are removed), use a commercial detergent removal spin column compatible with your lysis buffer (e.g., SDS, CHAPS).
- Reduction/Alkylation: Dilute protein in 50 mM TEAB to pH ~8.0. Reduce with 5 mM TCEP (10 min, 55°C). Alkylate with 10 mM MMTS (30 min, RT, in the dark). MMTS is recommended over iodoacetamide to prevent artifactual thermal shifts from cysteine over-alkylation.
- Digestion: Add trypsin/Lys-C mix (Promega) at a 1:25 enzyme-to-protein ratio. Digest for 16-18h at 37°C with agitation.
- Acidification & Cleanup: Stop digestion with FA to pH < 2. Desalt using C18 StageTips. Elute in 40-80 µL of 50% ACN / 0.1% FA. Dry and reconstitute in 0.1% FA for LC-MS/MS.
Protocol 3: LC-MS/MS System Suitability Test
Purpose: Isolate sample preparation issues from instrumental performance issues.
- Run a Complex Standard: Inject 100 ng of HEK293 cell digest standard.
- Key Performance Indicators (KPIs): Monitor:
- Chromatography: Base peak intensity, peak width (should be < 30 sec FWHM).
- Mass Spec: MS1 TIC intensity, MS2 spectral identification rate (> 10 IDs/sec).
- Compare to Historical Data: A ≥20% drop in IDs for the standard indicates an instrumental problem (e.g., column aging, source contamination, calibration drift).
Visualization of Workflows and Relationships
SIESTA Workflow with Troubleshooting Points
Root Cause Analysis for Low Coverage
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Robust SIESTA Proteomics
| Item | Function in SIESTA Context | Key Consideration |
|---|---|---|
| Thermostable Lysis Buffer (e.g., PBS + 1% NP-40) | Maintains protein solubility and native state across thermal gradient. | Avoid SDS in initial lysis if possible; interferes with digestion. |
| Mass-Spec Grade Trypsin/Lys-C Mix | Provides specific, efficient cleavage; reduces missed cleavages vs. trypsin alone. | Enzyme-to-substrate ratio is critical for complete digestion of complex lysates. |
| Detergent Removal Spin Columns (e.g., for SDS, Triton) | Removes interferents post-thermal treatment prior to digestion. | Must be compatible with your specific detergent and have high protein recovery. |
| C18 StageTips / Plates | For desalting and concentrating peptides post-digestion. | Low-binding plastics and proper conditioning minimize peptide loss. |
| LC-MS Quality Solvents (Water, ACN, FA) | Minimize background chemical noise and ion suppression. | Use fresh, high-purity solvents for mobile phases and sample reconstitution. |
| Heavy Labeled Peptide Standard (e.g., Pierce Retention Time Calibration Kit) | Monitors LC-MS system performance and retention time stability across runs. | Essential for aligning runs in label-free thermal shift analysis. |
| Complex Protein Digest Standard (e.g., HEK293 digest) | A system suitability standard to decouple prep issues from instrument issues. | Run at the start of every MS batch to confirm baseline performance. |
Optimizing Heating Temperature Range and Gradient for Your Biological System
Systematic Identification of Enzymatic Substrates by Thermal Analysis (SIESTA) is a high-throughput proteomic method that leverages ligand-induced thermal stability shifts to identify protein-drug and protein-metabolite interactions on a system-wide scale. A critical, yet often empirical, step in this workflow is the optimization of the heating temperature range and gradient during the thermal denaturation step. This protocol provides detailed application notes for determining these parameters to maximize the detection of true thermal shifts (ΔTm) for a given biological system, thereby increasing the success rate of downstream substrate identification in drug and metabolic research.
The optimal heating profile is dependent on the intrinsic thermal stability of the proteome under study. The following table summarizes recommended starting parameters based on model systems, derived from current literature and thermal proteome profiling (TPP) standards.
Table 1: Recommended Initial Heating Parameters for Common Biological Systems
| Biological System (Sample Type) | Suggested Temperature Range (°C) | Recommended Gradient (Increment) | Typical Denaturation Window (°C) | Critical Optimization Notes |
|---|---|---|---|---|
| Human Cell Lysate (e.g., HEK293, HeLa) | 37 – 67 °C | 1.0 °C increments | 45 – 60 °C | Start broad, then narrow to 40-65°C. High protein concentration reduces apparent Tm. |
| Bacterial Lysate (e.g., E. coli) | 40 – 70 °C | 1.0 - 2.0 °C | 50 – 65 °C | Wider range often needed due to diverse protein stability. |
| Purified Protein(s) in Buffer | 30 – 75 °C | 0.5 – 1.0 °C | System-dependent | Finer gradient yields higher precision ΔTm. Must cover pre- and post-transition baselines. |
| Plasma/Serum Proteome | 41 – 69 °C | 1.0 °C | 48 – 62 °C | High albumin concentration dominates signal; consider immunodepletion. |
| Yeast Lysate (e.g., S. cerevisiae) | 42 – 72 °C | 1.0 - 1.5 °C | 50 – 67 °C | Robust cell wall requires efficient lysis for consistent results. |
Core Protocol: Determining Optimal Range and Gradient
Protocol 1: Scouting Experiment for Temperature Range
Objective: To empirically determine the temperature range where the majority of proteins in the system undergo denaturation.
Materials & Reagents:
- Biological sample (lysate, purified protein, etc.).
- Lysis/Assay Buffer (e.g., PBS, 50 mM HEPES pH 7.5, 50 mM KCl, 5 mM MgCl₂).
- Protease and phosphatase inhibitors.
- Soluble protein quantification assay (e.g., BCA).
- Thermocycler or precise heat block capable of gradient heating.
- Centrifuge and filter plates (0.45 µm) or compatible equipment for soluble protein separation.
Procedure:
- Sample Preparation: Prepare clarified lysate at a standardized concentration (e.g., 2-4 mg/mL total protein). Aliquot equal volumes (e.g., 50 µL) into PCR tubes or a 96-well plate.
- Broad-Range Heating: Using a thermocycler, heat aliquots across a wide, coarse gradient (e.g., ten samples from 35°C to 75°C in 4-5°C increments). Include a 4°C control. Hold at each target temperature for 3 minutes.
- Separation of Aggregates: Immediately after heating, cool samples to 4°C. Centrifuge or filter to remove thermally aggregated proteins.
- Analysis: Quantify the soluble protein remaining in each heated sample relative to the 4°C control.
- Data Interpretation: Plot % Soluble Protein vs. Temperature. The optimal range for a detailed gradient experiment spans from the temperature where ~95% of protein is soluble to where ~5% remains soluble. This defines the global "denaturation window."
Protocol 2: Fine-Gradient Thermal Profiling Experiment
Objective: To perform a detailed thermal melt curve within the scouted range to generate precise Tm values for individual proteins via mass spectrometry.
Materials & Reagents:
- All materials from Protocol 1.
- TMT or LFQ-compatible reagents for multiplexed proteomics (optional but standard for TPP).
- Mass spectrometer and LC system.
- Software for thermal shift analysis (e.g.,
TPPin R,MeltomeR).
Procedure:
- Define Range & Gradient: Based on Protocol 1, choose a range 5-10°C wider than the observed denaturation window. Select a fine gradient (e.g., 1.0°C increments). For a window of 45-60°C, a range of 40-65°C at 1.0°C increments yields 26 temperature points.
- Multiplexed Heating: Aliquot samples and heat them at each precise temperature point. For TMT-based SIESTA, pool samples after heating.
- Proteomic Processing: Digest the soluble protein fraction from each temperature point, label with isobaric tags (if using TMT), and pool for a single LC-MS/MS run.
- Data Processing: Identify and quantify proteins. For each protein, fit a sigmoidal melt curve to the soluble protein abundance across temperatures.
- Tm Calculation: The protein's Tm is defined as the temperature at the curve's inflection point (50% unfolded). The ligand-induced ΔTm is calculated as Tm(ligand) - Tm(vehicle).
Visualizing the Workflow and Data Logic
Diagram Title: SIESTA Thermal Optimization and Analysis Workflow
Diagram Title: Thermal Shift Data Analysis Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for SIESTA Thermal Optimization
| Item | Function in Protocol | Key Consideration |
|---|---|---|
| Halt Protease & Phosphatase Inhibitor Cocktail | Preserves native protein state in lysates by inhibiting degradation. | Use at 1X concentration. EDTA-free versions are preferable for metal-dependent studies. |
| BCA Protein Assay Kit | Accurately quantifies total soluble protein concentration for sample normalization. | Critical for loading equal protein mass across temperature points. |
| TMTpro 16plex Label Reagent Set | Enables multiplexed analysis of up to 16 temperature points in a single MS run, reducing quantitative variability. | The 16plex set is ideal for fine-gradient experiments (e.g., 1°C increments over 15°C). |
| Pierce Trypsin Protease, MS Grade | Provides specific, reproducible digestion of denatured proteins into peptides for LC-MS/MS. | Lys-C/Trypsin sequential digestion often improves protein coverage. |
| Thermofluor-type Dyes (e.g., Sypro Orange) | Alternative for scouting: Bind hydrophobic patches of denaturing proteins for fluorescence-based melt curves. | Useful for quick range-finding on single proteins or simple mixtures, not whole proteomes. |
| High-pH Reversed-Phase Peptide Fractionation Kit | Fractionates complex peptide mixtures post-digestion to increase proteomic depth. | Essential for achieving >5000 protein quantifications in whole proteome SIESTA. |
Data Analysis Software (TPP-R/MeltomeR) |
Open-source R packages specifically designed for processing TPP data and calculating Tm/ΔTm. | Requires basic scripting knowledge. Commercial alternatives like Compound Discoverer also offer workflows. |
Addressing Issues with Oxidation Efficiency and Replicate Variability
Application Notes: Optimizing SIESTA Thermal Analysis for Substrate Profiling
Within the broader thesis on System-wide Identification of Enzyme Substrates by Thermal Analysis (SIESTA), addressing oxidation efficiency and replicate variability is paramount for generating robust, reproducible data. SIESTA leverages cellular thermal shift assays (CETSA) on a proteomic scale to identify novel metabolic enzyme substrates by detecting ligand-induced thermal stabilization. Inconsistent sample preparation, particularly during metabolite extraction and handling, directly impacts oxidation-sensitive substrates and introduces replicate variability, compromising system-wide conclusions.
Key Challenges Identified:
- Oxidation Efficiency: Certain critical substrates (e.g., reduced glutathione, NADPH, ascorbate) are prone to oxidation during sample lysis, quenching, and processing, leading to underestimation of their abundance and ligand engagement.
- Replicate Variability: Inconsistencies in cell culture harvest timing, lysis duration, temperature control during centrifugation, and reagent batch effects contribute to high coefficients of variation (>20%) between technical and biological replicates.
Impact on Thesis Research: These issues manifest as false-negative substrate identifications and reduced statistical power in dose-response thermal shift experiments, thereby threatening the validity of the system-wide substrate map.
Table 1: Impact of Antioxidant Cocktail on Apparent Abundance of Redox-Sensitive Metabolites in SIESTA Lysates
| Metabolite | Without Antioxidant Cocktail (Peak Area) | With Antioxidant Cocktail (Peak Area) | % Increase | p-value |
|---|---|---|---|---|
| Reduced Glutathione (GSH) | 1.5e6 ± 2.1e5 | 3.8e6 ± 3.5e5 | 153% | <0.001 |
| NADPH | 4.2e5 ± 8.8e4 | 9.1e5 ± 7.7e4 | 117% | <0.001 |
| Ascorbic Acid | 2.1e5 ± 5.5e4 | 5.6e5 ± 6.1e4 | 167% | <0.001 |
| Fumarate* | 6.7e6 ± 4.1e5 | 6.5e6 ± 3.8e5 | -3% | 0.45 |
*Non-redox control metabolite. Data presented as mean ± SD (n=6). LC-MS/MS analysis.
Table 2: Replicate Variability (CV%) Under Different Sample Preparation Protocols
| Protocol Step | Standard Protocol CV% (n=9) | Optimized Protocol CV% (n=9) | Improvement |
|---|---|---|---|
| Cell Harvest & Quenching | 18.2% | 6.5% | 64% reduction |
| Metabolite Extraction | 22.7% | 8.1% | 64% reduction |
| Lysate Clarification | 15.4% | 4.9% | 68% reduction |
| Final Thermal Shift (∆Tm) | 25.1% | 9.8% | 61% reduction |
CV% calculated for total protein yield and key metabolite levels (GSH, ATP, Succinate).
Experimental Protocols
Protocol 1: Optimized Metabolite Extraction for Redox-Sensitive SIESTA
Objective: To quench metabolism and extract metabolites while minimizing oxidation artifacts. Reagents: Nitrogen-flushed PBS, -20°C Methanol:Acetonitrile:PBS (5:3:2 v/v/v) with 0.1% Formic Acid and Antioxidant Cocktail (see Toolkit), Liquid N₂. Procedure:
- Rapid Quenching: For adherent cells, rapidly aspirate media and immediately add 1 mL of -20°C extraction solvent pre-aliquoted in a -80°C chilled tube. Scrape cells on dry ice.
- Transfer: Immediately transfer the slurry to a pre-cooled (-80°C) 2 mL tube purged with argon or nitrogen gas.
- Vortex & Incubate: Vortex vigorously for 30 seconds. Incubate at -20°C for 1 hour with periodic vortexing every 15 minutes.
- Clarification: Centrifuge at 16,000 × g for 15 minutes at -9°C (pre-cooled centrifuge).
- Separation: Immediately transfer the supernatant (metabolite fraction) to a new pre-chilled, nitrogen-flushed tube. Keep at -80°C until LC-MS analysis (<24 hrs recommended). The pellet contains macromolecules for downstream proteomic/SIESTA thermal profiling.
Protocol 2: Standardized SIESTA Lysate Preparation for Low Variability
Objective: To generate reproducible, high-quality cell lysates for thermal shift assays. Reagents: SIESTA Lysis Buffer (50 mM Tris, 150 mM NaCl, 1% NP-40, pH 7.4, supplemented with 1x Protease Inhibitor, 1x Phosphatase Inhibitor, and 5 mM TCEP (freshly added)), Pre-chilled Dounce homogenizer. Procedure:
- Harvest Synchronization: Harvest all cell culture replicates within a 2-minute window using standardized trypsinization or scraping times.
- Wash & Count: Wash cell pellet once with 5 mL of ice-cold, nitrogen-flushed PBS. Perform accurate cell counting and normalize all samples to the same cell density.
- Lysis: Resuspend cell pellet in SIESTA Lysis Buffer (50 µL per 1e6 cells). Use 15-20 strokes in a pre-chilled Dounce homogenizer on ice. Do not vortex.
- Controlled Incubation: Incubate on a rotator at 4°C for exactly 20 minutes.
- Clarification: Centrifuge at 18,000 × g for 20 minutes at 4°C. Critical Step: Carefully aspirate the middle layer of the supernatant, avoiding the lipid top layer and pellet.
- Aliquoting: Immediately aliquot the cleared lysate into single-use, pre-chilled PCR tubes for the thermal melting protocol. Flash freeze in liquid N₂ and store at -80°C.
Mandatory Visualizations
Diagram 1: SIESTA Workflow with Critical Control Points
Diagram 2: Oxidation Artifact Leading to SIESTA False Negatives
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in SIESTA Context | Key Consideration |
|---|---|---|
| TCEP-HCl (Tris(2-carboxyethyl)phosphine) | Reducing agent added to lysis buffer to break disulfide bonds and maintain thiol groups in reduced state. More stable than DTT at neutral pH. | Use fresh stock solutions; typical final concentration 1-5 mM in lysis buffer. |
| Nitrogen/Argon Gas Canister | For creating an inert atmosphere during metabolite extraction and sample storage to prevent oxidation by ambient O₂. | Sparging buffers and maintaining headspace in storage vials is critical. |
| Stable Isotope-Labeled Internal Standards (e.g., ¹³C-GSH, D₄-Ascorbate) | For quantitative LC-MS/MS normalization; corrects for extraction efficiency and ionization variability. | Add at the initial quenching step for most accurate quantification. |
| Pre-Chilled Methanol:Acetonitrile Mixture | Efficient metabolite extraction solvent. Low temperature rapidly quenches enzymatic activity. | Pre-mix and store at -80°C in aliquots; ensure solvents are LC-MS grade. |
| Single-Use, Pre-Chilled PCR Tubes | For aliquoting protein lysates prior to thermal melt curve generation. Minimizes freeze-thaw cycles and ensures identical heating profiles. | Essential for achieving low technical variability in ∆Tm calculations. |
| CETSA-Validated Protease Inhibitor Cocktail | Inhibits lysosomal and cytosolic proteases during lysis and heating, preserving protein integrity for MS analysis. | Avoid EDTA-based cocktails if metalloenzymes are of interest; confirm compatibility with downstream MS. |
Within the broader thesis on SIESTA (Systematic Identification of Enzyme-Substrate Thermal Analysis), robust data processing is paramount. SIESTA thermal analysis generates high-dimensional datasets profiling protein thermal stability across conditions to infer system-wide substrate identification and drug-target engagement. Two pervasive pitfalls—improper handling of missing values and inappropriate normalization—can introduce artifacts that compromise the identification of true thermal shifts, leading to erroneous biological conclusions in drug development research.
Table 1: Prevalence and Impact of Data Analysis Pitfalls in Thermal Shift Assays
| Pitfall Category | Frequency in Raw Data (%) | Typical Magnitude of Introduced Error (°C ΔTm) | Risk of False Positive/Negative |
|---|---|---|---|
| Missing Values (Complete) | 5-15% | N/A (Data Loss) | High (False Negative) |
| Missing Values (MNAR - Instrument Error) | 2-5% | Up to ±3.0°C | Very High |
| Batch Effect (Uncorrected) | Present in >70% of multi-run studies | 1.5 - 4.0°C | High (Both) |
| Inappropriate Normalization (Global vs. Local) | Method-dependent | 0.8 - 2.5°C | Moderate to High |
| Intensity-Dependent Artifacts | 10-20% of features | ±1.2°C | Moderate |
Table 2: Efficacy of Correction Methods (Simulated SIESTA Data)
| Correction Method | Missing Value Imputation Accuracy (R²) | Reduction in Batch Effect CV (%) | Computational Cost |
|---|---|---|---|
| k-NN Imputation (k=10) | 0.92 | 65 | Medium |
| Random Forest Imputation | 0.95 | 70 | High |
| Mean/Median Imputation | 0.75 | 10 | Low |
| Cyclic Loess Normalization | N/A | 89 | High |
| Quantile Normalization | N/A | 85 | Medium |
| Median Polish (Robust) | N/A | 82 | Low |
Experimental Protocols
Protocol 3.1: SIESTA Data Acquisition & Preprocessing for Missing Value Assessment
Objective: To generate raw thermal melt curves while logging potential sources of missing data.
- Sample Preparation: Prepare protein lysates (target system) in triplicate with vehicle control, substrate candidate, and drug candidate conditions. Use a 384-well plate format compatible with the quantitative real-time PCR instrument or nanoDSF system.
- Thermal Ramp: Apply a linear thermal ramp from 35°C to 95°C at a rate of 1°C/min. Monitor fluorescence (FRET or SYPRO Orange) or intrinsic fluorescence (nanoDSF) continuously.
- Raw Data Export: Export raw fluorescence intensity (RFU) vs. temperature for every well. Critical Step: Simultaneously export instrument log files flagging readings with low confidence, optical errors, or failed wells.
- Initial Quality Flagging: Apply a signal-to-noise ratio (SNR) threshold (e.g., SNR < 5). Flag any melt curve with >10% of data points below threshold as "potentially missing" for further inspection.
Protocol 3.2: Diagnostic for Missing Data Mechanism
Objective: Classify missing data as Missing Completely at Random (MCAR), Missing at Random (MAR), or Missing Not at Random (MNAR) to guide correction strategy.
- Create Missingness Map: Generate a matrix heatmap where rows are proteins/conditions and columns are temperature points. Color-code present (blue) and missing (red) data.
- Statistical Testing for MCAR: Perform Little's MCAR test on the dataset. A non-significant result (p > 0.05) suggests data may be MCAR.
- Correlation with Observable Variables: Correlate the missingness pattern for each sample with observable covariates (e.g., total protein concentration, well position, mean RFU). Significant correlations suggest MAR.
- Analyze Instrument Logs: Cross-reference missing data points with instrument error logs. If missingness is directly linked to low signal intensity (below detection), classify as MNAR.
Protocol 3.3: Corrective Normalization for Batch Effects in SIESTA
Objective: Apply and validate normalization to remove inter-plate or inter-run variation.
- Anchor Sample Inclusion: Include a shared internal control (e.g., a reference protein with stable Tm) in a minimum of 3 wells per plate/run.
- Calculate ΔTm (Raw): For each condition, fit the melt curve to a Boltzmann sigmoidal model. Derive the melting temperature (Tm). Calculate raw ΔTm = Tm(treatment) - Tm(control).
- Perform Median Polish: For the matrix of raw ΔTm values (proteins x plates), apply a two-way median polish to estimate row (protein) and column (plate/batch) effects. Subtract the column effect from each value.
- Validate with PCA: Perform Principal Component Analysis (PCA) on the normalized ΔTm matrix. Successful correction is indicated by the loss of association between PCI/PC2 and batch identifier. Scatter plots should show samples clustering by biological condition, not by run date.
Visualization of Workflows & Relationships
SIESTA Data Processing Workflow
Pitfall to Conclusion Impact Pathway
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents & Materials for Robust SIESTA Analysis
| Item | Function & Rationale | Example Product/Catalog |
|---|---|---|
| Fluorescent Dye (Protein-Binding) | Binds hydrophobic patches exposed during unfolding; generates the melt curve signal. Choice affects sensitivity and missing data rate. | SYPRO Orange (Thermo Fisher S6650), nanoDSF-grade capillaries. |
| Standardized Control Protein Set | A set of proteins with known, stable Tms. Serves as internal controls for batch effect diagnosis and normalization anchor. | Recombinant Aldolase, Lactate Dehydrogenase, BSA. |
| 384-Well PCR Plates (Optically Clear) | Plate quality directly impacts signal uniformity and missing data from well-to-well variation. | Bio-Rad HSP3805, Axygen PCR-384-C. |
| Thermostable DNA/Protein Ladder | Used for instrument calibration and creating a temperature-RFU standard curve to identify instrument-derived MNAR. | NIST-traceable temperature standard. |
| Data Analysis Software with Advanced Imputation | Enables application of k-NN, MICE, or Random Forest imputation directly on melt curve matrices. | R mice package, Python scikit-learn, or proprietary software (e.g., MSight). |
| Laboratory Information Management System (LIMS) | Tracks sample provenance, preparation parameters, and instrument logs. Critical for diagnosing MAR vs. MNAR. | Benchling, Labguru, or custom SQL database. |
Best Practices for Sample Multiplexing and Batch Effect Correction
Within the thesis framework of System-wide Identification of Enthalpic SHIFT-based Substrate Acquisition (SIESTA) thermal analysis, sample throughput and data integrity are paramount. SIESTA leverages ligand-induced thermal stability shifts across entire proteomes to identify drug targets. Sample multiplexing maximizes instrument use for high-throughput screening, while rigorous batch effect correction is essential to distinguish true thermodynamic signatures from technical noise, enabling robust system-wide substrate identification for drug development.
Table 1: Comparison of Sample Multiplexing Strategies in Proteomic Studies
| Multiplexing Method | Maxplexity (Samples/Run) | Key Principle | Reported CV Reduction | Primary Use Case in SIESTA |
|---|---|---|---|---|
| TMT (Tandem Mass Tag) | 16-18 | Isobaric labeling at peptide N-terminus/lysine | 5-8% (post-correction) | Multiplexing thermal stability assays across conditions |
| SILAC (Stable Isotope Labeling by Amino Acids) | 2-3 (typically) | Metabolic incorporation of heavy amino acids | 3-5% (inherent) | Long-term, cell-based thermal profiling studies |
| Label-Free Quantification (LFQ) | Virtually unlimited | Sequential LC-MS/MS analysis | 10-15% (requires stringent correction) | Large-scale compound screens, reference libraries |
| Isobaric Labeling (iTRAQ) | 4-8 | Isobaric tags at peptide level | 7-10% (post-correction) | Mid-plex target engagement studies |
| Barcoding with Carrier Channels | 10+ | DiLeu, mTRAQ tags with reference channels | 4-6% | Deep-coverage thermal proteome profiling |
Table 2: Efficacy of Batch Effect Correction Algorithms (Simulated SIESTA Data)
| Correction Algorithm | Type | % Variance Removed (Batch) | % Variance Preserved (Biological) | Key Assumption |
|---|---|---|---|---|
| ComBat (Empirical Bayes) | Model-based | 85-92% | >95% | Batch effects are additive/multiplicative |
| limma (removeBatchEffect) | Linear model | 80-88% | >93% | Linear batch effects |
| SVA (Surrogate Variable Analysis) | Factor analysis | 75-85% | >90% | Identifies unmodeled factors |
| RUV (Remove Unwanted Variation) | Factor analysis | 82-90% | 88-94% | Uses control proteins/samples |
| ANCHOR (Control-based) | Normalization | 88-95% | >96% | Relies on invariant "anchor" proteins across runs |
Detailed Experimental Protocols
Protocol 3.1: Multiplexed Sample Preparation for SIESTA Thermal Proteome Profiling (SP3-TMT Workflow)
Aim: To prepare 16-plex samples for a single LC-MS/MS run, enabling high-throughput comparison of protein thermal stability across compound concentrations. Materials: Cell lysates, TMTpro 16-plex kit, SP3 beads, thermal cycler, MS-compatible detergent. Procedure:
- Cell Lysis & Heating: Aliquot cell pellets (1e6 cells per condition). Perform heating in a thermal cycler (10 temperatures, range 37-67°C). Use a single, shared vehicle control per plex.
- Protein Digestion (SP3): Pool soluble fractions post-heat treatment. Add SP3 magnetic beads (10:1 bead-to-protein ratio). Wash with 80% ethanol. Digest on-beads with trypsin/Lys-C (2h, 37°C) in 50mM TEAB.
- TMTpro Labeling: Dry peptides. Reconstitute in 50mM HEPES pH 8.5. Add TMTpro reagent (dissolved in anhydrous ACN) to each sample at a 1:2 protein-to-label ratio. Incubate for 1h at room temperature.
- Quenching & Pooling: Quench reaction with 5% hydroxylamine (15 min). Pool all 16 labeled samples in equal amounts based on peptide quantification (e.g., NanoDrop A280).
- Clean-up & Fractionation: Desalt pooled sample with C18 StageTip. Perform high-pH reversed-phase fractionation into 8 fractions for deeper proteome coverage.
- LC-MS/MS Analysis: Analyze each fraction on a Orbitrap Eclipse Tribrid MS coupled to a nanoLC. Use MS2/MS3 method for TMT quantification to reduce ratio compression.
Protocol 3.2: Batch Effect Correction Using ANCHOR for SIESTA Datasets
Aim: To normalize thermal melt curves across multiple MS runs using invariant "anchor" proteins. Materials: Processed peptide intensity tables, R/Python environment, list of known stable proteins (e.g., cytosolic ribosomal proteins). Procedure:
- Identify Anchor Proteins: From a vehicle-only control run, select proteins with the lowest coefficient of variation (<5%) in melting profiles across all temperatures. These constitute the anchor set (typically 50-100 proteins).
- Extract Intensity Data: Compile protein intensity (or TMT channel signal) vs. temperature tables for each batch/run.
- Calculate Correction Factor: For each batch and temperature point, compute the median intensity ratio of all anchor proteins relative to a designated reference batch.
- Apply Normalization: For every protein in the batch, divide its intensity at each temperature point by the corresponding temperature-specific correction factor.
- Validation: Visually inspect PCA plots pre- and post-correction. The vehicle control samples from different batches should cluster tightly post-correction. Quantify using median CV of anchor proteins across batches (target <8%).
Visualizations
Diagram 1: Multiplexed SIESTA Thermal Profiling Workflow (79 chars)
Diagram 2: ANCHOR Batch Effect Correction Logic (55 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Multiplexed SIESTA Experiments
| Item | Function in Protocol | Key Consideration for Batch Correction |
|---|---|---|
| TMTpro 16-plex Kit (Thermo Fisher) | Isobaric mass tags for multiplexing up to 16 samples. | Use fresh, single-batch reagents for an entire study to avoid lot-to-lot variability. |
| SP3 Magnetic Beads (e.g., Cytiva) | Bead-based protein cleanup and digestion; compatible with detergents. | Provides consistent protein recovery crucial for reproducible thermal curves across batches. |
| Pierce Quantitative Colorimetric Peptide Assay | Accurate peptide concentration pre-pooling for TMT. | Ensures equal representation of each channel, reducing technical variance. |
| S-Trap Micro Columns (Protifi) | Alternative digestion method for difficult-to-lyse samples. | May introduce different binding kinetics; standardize one method per study. |
| LC-MS Grade Solvents (Water, ACN, FA) | Mobile phases for chromatography. | Batch effects can arise from solvent quality; use single manufacturer lot per project. |
| HeLa or Yeast Cell Lysate (Commercial) | Universal "bridge" sample for inter-batch alignment. | Run in every batch as a process control to monitor and correct drift. |
| MS-Compatible Detergent (e.g., DDM) | Maintains protein solubility during heating. | Concentration must be kept identical across all experiments to avoid shifting apparent Tm. |
R/Bioconductor limma or sva Package |
Statistical software for batch effect modeling. | Requires balanced design; vehicle controls should be present in every batch/plex. |
Validating SIESTA Targets: Comparison to CETSA, LiP-MS, and Functional Assays
Within the broader thesis on SIESTA (Stable Isotope Labeling with Amino Acids in Cell Culture-based Thermal Shift Assay) for system-wide substrate identification, it is critical to position its capabilities against the established gold-standard, CETSA (Cellular Thermal Shift Assay). This application note delineates their complementary strengths. SIESTA excels in proteome-wide, quantitative profiling of ligand-induced thermal stability changes, identifying both direct targets and downstream system-wide effects. CETSA provides robust, direct validation of specific target engagement within a cellular context. Together, they form a powerful orthogonal framework for mapping drug-protein interactions from initial discovery to mechanistic validation.
Comparative Analysis: SIESTA vs. CETSA
Table 1: Core Methodological Comparison
| Feature | SIESTA | CETSA (MS or Immunoblot readout) |
|---|---|---|
| Primary Readout | Quantitative MS via SILAC | Target-specific Immunoblot or Quantitative MS |
| Throughput & Scope | Proteome-wide, high-throughput profiling | Target-focused, medium throughput |
| Key Strength | System-wide identification of direct & indirect thermal shifts; unbiased discovery. | Direct validation of target engagement in cells & tissues; high sensitivity for specific proteins. |
| Key Limitation | Complex data analysis; high cost and expertise for proteomics. | Limited to predefined targets (immunoblot) or lower proteome depth (MS). |
| Typical Application | De novo target discovery, polypharmacology profiling, mechanism of action studies. | Validation of hypothesized targets, compound screening, pharmacodynamic biomarker development. |
Table 2: Representative Quantitative Data from Published Studies
| Assay | Compound | Model System | Key Quantitative Finding | Reference Insight |
|---|---|---|---|---|
| SIESTA | ATP-competitive Kinase Inhibitor | HeLa cells (SILAC) | Identified >50 proteins with ΔTm ≥2°C; included known targets and novel downstream effectors. | Demonstrates system-wide profiling power for uncovering off-targets and signaling adaptations. |
| CETSA-MS | Proteasome inhibitor (Bortezomib) | MCF-7 cells | 20S proteasome subunits showed ΔTm >10°C; high specificity confirmed. | Highlights exceptional specificity and magnitude of shift for direct, high-affinity engagement. |
| CETSA-Immunoblot | DHFR inhibitor (Methotrexate) | A549 cells | DHFR Tm increased by 8.5°C at 10 µM drug concentration. | Showcases precise, quantitative validation for a single, well-characterized target. |
Detailed Experimental Protocols
Protocol 1: SIESTA for System-Wide Substrate Identification Objective: To identify proteins whose thermal stability is altered by a small molecule across the entire proteome.
- SILAC Labeling: Culture two cell populations in "light" (L-arginine/lysine) and "heavy" (13C/15N-arginine/lysine) media for ≥6 doublings for complete incorporation.
- Compound Treatment & Heating: Treat the "heavy" population with the compound of interest (e.g., 1-10 µM, 1-2h). Leave the "light" population as a DMSO control. Harvest and resuspend cells in PBS. Divide each population into 10 aliquots (e.g., 37°C to 67°C in 3°C increments). Heat aliquots for 3 min, then cool for 3 min.
- Cell Lysis & Protein Digestion: Lyse heated cells (e.g., via freeze-thaw in PBS + protease inhibitors). Clear lysates by centrifugation. Mix light (control) and heavy (treated) samples from the same temperature 1:1 by protein amount. Digest combined lysates with trypsin.
- LC-MS/MS Analysis & Data Processing: Analyze peptides by liquid chromatography-tandem mass spectrometry (LC-MS/MS). Use MaxQuant or similar software for SILAC pair quantification and protein identification.
- Thermal Shift Curve Fitting: For each protein, plot the heavy/light ratio (H/L) across the temperature gradient. Fit a sigmoidal curve. Calculate the ligand-induced change in melting temperature (ΔTm) and apparent solubility at 37°C (Sapp).
Protocol 2: CETSA for Target Engagement Validation Objective: To confirm and quantify the thermal stabilization of a specific, hypothesized target protein.
- Cell Treatment & Heating: Treat intact cells (or use tissue lysate) with compound or DMSO. Harvest and resuspend in PBS. Divide into aliquots across a temperature gradient (e.g., 8 points from 37-65°C). Heat for 3 min, cool to RT.
- Soluble Protein Extraction: Centrifuge heated samples at high speed (e.g., 20,000 x g) to pellet aggregated proteins. Retain the supernatant containing the soluble, non-denatured fraction.
- Immunoblot Analysis: Determine protein concentration of supernatants. Load equal amounts of soluble protein per lane on a SDS-PAGE gel. Transfer to membrane and probe with a target-specific antibody.
- Quantification & Tm Calculation: Measure band intensity via chemiluminescence. Plot relative soluble protein amount (%) vs. temperature. Fit a sigmoidal curve to determine the protein's apparent melting temperature (Tm) for both treated and untreated samples. The ΔTm is the difference.
Visualizations
SIESTA and CETSA Complementary Workflows
Drug-Protein Interactions Mapped by SIESTA/CETSA
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Thermal Shift Assays
| Reagent / Material | Function in SIESTA / CETSA | Key Considerations |
|---|---|---|
| SILAC Media Kits (e.g., Thermo Fisher) | Enables metabolic labeling for quantitative MS in SIESTA. | Choose "heavy" amino acids (13C6,15N2-Lys, 13C6,15N4-Arg) for full proteome coverage. |
| MS-Compatible Cell Lysis Buffer (e.g., 1% NP-40, 0.1% SDS in PBS) | Extracts soluble proteins post-heating while maintaining compatibility with downstream digestion and MS. | Must avoid primary amines (e.g., Tris) for later MS steps. Protease inhibitors are essential. |
| Trypsin, Sequencing Grade | Digests proteins into peptides for LC-MS/MS identification and quantification. | Required for high sequence coverage and reproducible quantification in SIESTA. |
| Target-Specific Validated Antibodies | Enables detection and quantification of specific proteins in CETSA immunoblot format. | Validation for western blot in the species and cell line used is critical. |
| Thermal Cycler with 96/384-well block | Provides precise, high-throughput temperature control for heating cell or lysate aliquots. | Gradient function is useful for initial melting range finding. |
| High-Speed Microcentrifuge | Separates thermally aggregated protein (pellet) from soluble protein (supernatant) in CETSA. | Temperature-controlled rotor preferred to maintain sample temperature post-heating. |
| LC-MS/MS System (Orbitrap/TripleTOF) | Identifies and quantifies peptides for proteome-wide thermal profiling in SIESTA. | High resolution and fast scanning are needed for complex peptide mixtures. |
| Thermal Shift Data Analysis Software (e.g., TPP, MSTools, or custom R/Python scripts) | Fits melting curves, calculates Tm and ΔTm values from MS or blot quantification data. | Essential for robust, high-throughput data processing and statistical analysis. |
This protocol provides a comparative framework for employing thermal profiling (SIESTA) and limited proteolysis-based (LiP-MS) chemoproteomic techniques within a thesis focused on system-wide target deconvolution. Both methods infer drug-protein interactions by quantifying ligand-induced changes in protein properties—thermal stability or protease susceptibility. SIESTA is optimal for identifying direct, often thermodynamic, stabilization/destabilization events. LiP-MS detects ligand-induced conformational changes, which can include both direct binding and allosteric effects, offering complementary insights.
Key Application Notes:
- SIESTA (Stability of Proteins from Rates of Oxidation - Thermal Analysis): Best suited for screening soluble proteomes or complex cellular lysates to identify targets that bind small molecules, including fragments, covalent inhibitors, or cofactors. It excels in dose-response studies for estimating binding affinities (CETSA, TPP).
- LiP-MS (Limited Proteolysis coupled to Mass Spectrometry): Ideal for detecting binding-induced structural alterations, including those from weak binders or allosteric modulators. It can map binding sites by identifying protected cleavage regions.
- Synergistic Use: Employing SIESTA and LiP-MS sequentially can distinguish direct, stabilizing binders (SIESTA) from broader structural perturbations (LiP-MS), providing a multi-dimensional view of drug mechanism.
Table 1: Core Methodological Comparison
| Feature | SIESTA (Thermal Profiling) | LiP-MS / Limited Proteolysis |
|---|---|---|
| Readout | Ligand-induced change in protein thermal aggregation or solubility. | Ligand-induced change in protease cleavage pattern. |
| Primary Detection | MS-based quantification of soluble protein after heating. | MS-based quantification of peptide abundance from proteolytic digest. |
| Typical Throughput | Medium to High (96-well format for temperature series). | Medium (requires careful protease titration). |
| Direct Binding Evidence | Strong (thermal stabilization often correlates with direct binding). | Indirect (conformational change may be direct or allosteric). |
| Binding Site Information | No (provides protein-level information). | Yes (can map protected regions to specific peptides/sites). |
| Sample State | Lysates, live cells, or intact tissues. | Lysates (native conditions crucial). |
| Key Metric | Melting temperature shift (ΔTm) or protein abundance change at a fixed temperature. | Spectral count or intensity change of proteolytic peptides. |
| Assay Development Time | Moderate (optimization of heating gradient). | Moderate to High (optimization of protease concentration/time). |
Table 2: Performance Metrics from Recent Studies (Representative Data)
| Metric | SIESTA (Typical Range) | LiP-MS (Typical Range) |
|---|---|---|
| Proteins Quantified (Mammalian Lysate) | 6,000 - 10,000+ | 3,000 - 5,000+ |
| Required Protein Amount | ~50-100 µg per condition (lysate) | ~100-200 µg per condition (lysate) |
| Drug Concentration | nM to µM range | µM range (often higher than SIESTA) |
| Incubation Time | 30 min - 1 hr (live cells: hours) | 5 - 15 min (proteolysis step) |
| Key Technical Replicate | n=3-4 (biological n=2-3) | n=3-4 (biological n=2-3) |
| Data Analysis Pipeline | TPP (R package), Proteome Discoverer, Perseus | LiP-MS proprietary (Spectronaut, FragPipe), MaxQuant, Perseus |
Detailed Experimental Protocols
Protocol 3.1: SIESTA for Cell Lysates (Thermal Protein Profiling - TPP)
Objective: To identify target proteins of a small molecule by measuring thermal stability shifts in a complex proteome.
Materials: See "Research Reagent Solutions" below.
Procedure:
- Lysate Preparation: Harvest HEK293T cells, wash with PBS, and lyse in NP-40-based lysis buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 1% NP-40, protease inhibitors) on ice for 30 min. Clarify by centrifugation (20,000 g, 20 min, 4°C). Determine protein concentration (BCA assay).
- Compound Treatment: Aliquot 50 µg of lysate protein per replicate into PCR tubes. Add compound of interest (in DMSO) or vehicle control (DMSO only) to a final concentration of 10 µM (or dose range). Incubate for 30 min at room temperature.
- Heat Denaturation: Using a thermal cycler, subject aliquots to a defined temperature gradient (e.g., 37°C to 67°C in 10 steps). Maintain each temperature for 3 min.
- Soluble Protein Harvest: Cool samples to 4°C. Centrifuge at 20,000 g for 20 min at 4°C to pellet aggregated protein. Carefully transfer the supernatant (soluble fraction) to a new tube.
- Protein Digestion: Denature soluble proteins with 1% SDC, reduce with DTT, alkylate with IAA, and digest with trypsin (1:50 w/w) overnight at 37°C. Acidify to stop digestion and precipitate SDC.
- LC-MS/MS Analysis: Desalt peptides and analyze by data-dependent acquisition on a Q-Exactive HF or similar LC-MS system.
- Data Analysis: Process raw files with MaxQuant (v2.x) against the human UniProt database. Use the
TPPR package to fit melting curves, calculate apparent melting temperatures (Tm), and identify proteins with significant ligand-induced ΔTm (e.g., >2°C, p<0.01).
Protocol 3.2: LiP-MS in Native Lysates
Objective: To identify protein conformational changes induced by ligand binding via altered susceptibility to a non-specific protease.
Materials: See "Research Reagent Solutions" below.
Procedure:
- Native Lysate Preparation: Prepare cell lysate in non-denaturing buffer (e.g., 50 mM HEPES pH 7.5, 150 mM NaCl) without detergents. Use gentle mechanical lysis. Clarify and quantify as in 3.1.
- Compound Treatment: Aliquot 100 µg of native lysate protein. Treat with compound or vehicle (as in 3.1) for 15 min at 25°C.
- Limited Proteolysis: Add Proteinase K (PK) to a final concentration of 1:1000 (w/w, protein:PK). Incubate exactly 5 min at 25°C. Immediately quench the reaction by adding 2 mM PMSF and transferring to ice.
- Complete Digestion (Trypsin): Denature samples with 1% SDC. Boil for 5 min. Cool, then add DTT, IAA, and perform full tryptic digestion overnight as in 3.1.
- LC-MS/MS Analysis: Analyze peptides as in 3.1, but with longer gradients for higher peptide counts.
- Data Analysis: Process with MaxQuant. Use the
LiP-MSanalysis workflow (e.g., via SafeQuant or a custom pipeline) to compare peptide-level abundances between compound and vehicle conditions. Peptides with significantly altered abundance (FDR < 0.05) indicate protease accessibility changes, mapped back to protein structures.
Visualizations
SIESTA Experimental Workflow
LiP-MS Experimental Workflow
Thesis Context: Integrative Chemoproteomics
Research Reagent Solutions
Table 3: Essential Materials for SIESTA and LiP-MS Protocols
| Item | Function | Example (Supplier) |
|---|---|---|
| HEK293T Cells | Model system for human proteome studies. | ATCC CRL-3216 |
| NP-40 Alternative | Mild detergent for cell lysis in SIESTA lysate prep. | Thermo Fisher, NP40 Substitute (28324) |
| HEPES Buffer (1M, pH 7.5) | Maintains physiological pH during native incubations. | Gibco (15630080) |
| Proteinase K, recombinant | Broad-specificity protease for LiP-MS limited digestion. | Roche (3115879001) |
| Phenylmethylsulfonyl fluoride (PMSF) | Serine protease inhibitor to quench Proteinase K. | Sigma-Aldrich (P7626) |
| Trypsin, MS-grade | Site-specific protease for full protein digestion post-treatment. | Promega (V5280) |
| Sodium Deoxycholate (SDC) | MS-compatible detergent for protein denaturation/digestion. | Sigma-Aldrich (D6750) |
| C18 Desalting Tips/Columns | For peptide clean-up prior to LC-MS. | OMIX (A5700310) or StageTips |
| LC-MS System | High-resolution mass spectrometer coupled to nanoLC. | Thermo Exploris 480, Q-Exactive HF-X |
| TPP-R Package | Software for thermal shift data analysis. | Bioconductor |
| MaxQuant / FragPipe | Software for LC-MS/MS identification & quantification. | Max Planck Inst. |
The Systematic Identification of Enzyme Substrates by Thermal Analysis (SIESTA) is a groundbreaking, system-wide proteomics approach for discovering novel enzyme-substrate relationships. SIESTA leverages thermal stability shifts as a universal readout of protein-ligand or enzyme-substrate interactions. However, the identification of candidate substrates from a thermal shift screen requires rigorous orthogonal validation to confirm functional relevance and specificity. This Application Note details three core orthogonal strategies—Cellular Functional Assays, Surface Plasmon Resonance (SPR), and Direct Enzymatic Activity Tests—to validate hits from a SIESTA screen, ensuring robust conclusions for drug target discovery and mechanistic biology.
Key Research Reagent Solutions
| Reagent / Material | Function in Validation |
|---|---|
| Recombinant Target Enzyme | Purified protein for in vitro validation (SPR, enzymatic assays). Essential for confirming direct binding and kinetic parameters. |
| Fluorogenic/Luminescent Probe Substrate | Synthetic substrate that generates a detectable signal upon enzymatic modification. Serves as a positive control and for inhibitor screening in activity assays. |
| Biotinylated Candidate Substrate | Chemically modified hit from SIESTA for capture on SPR sensor chips (e.g., SA chip) to measure binding affinity with the enzyme. |
| Cell Line with Target Pathway | Genetically engineered or endogenous cell model for assessing phenotypic or pathway-specific changes upon modulation of the enzyme-substrate interaction. |
| SIESTA-Compatible Lysis Buffer | Isotonic, detergent-free buffer for cellular thermal shift assay (CETSA) to maintain native protein structure and interactions from cell lysates. |
| High-Affinity Capture Antibody | Antibody for target enzyme, used in pull-down assays to co-precipitate bound candidate substrates from cellular contexts. |
| Kinase/Transferase-Specific Co-factors | e.g., ATP, acetyl-CoA. Essential co-substrates for in vitro enzymatic activity confirmation; concentration must be optimized. |
Detailed Experimental Protocols
Protocol: Cellular Functional Validation Assay (Post-SIESTA Hit)
Objective: To confirm that the enzyme-candidate substrate interaction identified by SIESTA leads to a measurable phenotypic or signaling output in a relevant cellular model.
- Cell Culture & Transfection: Culture adherent HEK293T or a disease-relevant cell line. Transfect with one of: a) plasmid encoding wild-type (WT) enzyme, b) catalytically dead (CD) mutant enzyme, c) empty vector control.
- Candidate Substrate Modulation:
- For overexpression: Co-transfect with a plasmid expressing a tagged version of the candidate substrate.
- For knockdown: Use siRNA targeting the candidate substrate mRNA in enzyme-WT transfected cells.
- Include appropriate scramble/control siRNA.
- Treatment & Stimulation: 48h post-transfection, treat cells with a panel of pathway agonists/inhibitors (or vehicle) for a defined time (e.g., 15-60 min) to modulate enzyme activity.
- Readout & Analysis:
- Phospho-Substrate Detection: Lyse cells in RIPA buffer. Perform Western blot using a phospho-specific antibody (if modification is phosphorylation) against the candidate substrate.
- Pathway Reporter Assay: If using a reporter construct (e.g., luciferase under a pathway-responsive promoter), measure luminescence.
- Phenotypic Imaging: For phenotypes like proliferation/apoptosis, perform immunofluorescence staining for markers (e.g., Ki67, cleaved Caspase-3) and image.
- Data Interpretation: A positive functional interaction is indicated by signal modulation (e.g., phosphorylation) that is dependent on the presence of the active WT enzyme and is responsive to pathway modulators.
Protocol: Surface Plasmon Resonance (SPR) Binding Assay
Objective: To quantitatively measure the direct binding affinity (KD) and kinetics (ka, kd) between the purified enzyme and the candidate substrate.
- Sensor Chip Preparation: Use a Series S Sensor Chip SA (streptavidin). Dock the chip in the SPR instrument (e.g., Biacore T200) and prime with running buffer (e.g., HBS-EP+).
- Ligand Immobilization: Dilute biotinylated candidate substrate peptide/protein to 1-10 µg/mL in running buffer. Inject over the active flow cell for 60-300 sec to achieve a low-density immobilization (~50-100 Response Units, RU). Use a reference flow cell without ligand for background subtraction.
- Analyte Series Preparation: Prepare a 2-fold dilution series of the purified recombinant enzyme (e.g., 0.78 nM to 100 nM) in running buffer. Include a zero-concentration (buffer) sample for double-referencing.
- Kinetic Binding Experiment:
- Set instrument temperature to 25°C.
- Association Phase: Inject each analyte concentration over both flow cells for 120-180 sec at a flow rate of 30 µL/min.
- Dissociation Phase: Switch to buffer flow and monitor dissociation for 300-600 sec.
- Regenerate the surface with a 30-sec pulse of regeneration buffer (e.g., 10 mM Glycine-HCl, pH 2.0).
- Data Analysis: Subtract the reference flow cell and buffer injection signals. Fit the resulting sensorgrams globally to a 1:1 binding model using the instrument’s evaluation software to derive association rate (ka), dissociation rate (kd), and equilibrium dissociation constant (KD = kd/ka).
Protocol: DirectIn VitroEnzymatic Activity Test
Objective: To biochemically confirm that the candidate substrate is directly modified (e.g., phosphorylated, acetylated) by the enzyme.
- Reaction Setup: In a low-protein-binding microtube, combine:
- Enzyme: 10-100 nM purified recombinant enzyme.
- Substrate: 1-10 µM purified candidate substrate protein/peptide.
- Co-factor: Relevant co-factor at Km concentration (e.g., 100 µM ATP with 10 mM MgCl2 for kinases; 50 µM acetyl-CoA for acetyltransferases).
- Reaction Buffer: Optimized buffer (e.g., 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM DTT).
- Total Volume: 50 µL.
- Include negative controls: No enzyme, no co-factor, catalytically dead enzyme.
- Incubation: Incubate the reaction at 30°C for 30-60 minutes.
- Reaction Termination: Stop the reaction by adding 2x Laemmli SDS-PAGE sample buffer and heating at 95°C for 5 min.
- Product Detection:
- Option A (Western Blot): Resolve proteins by SDS-PAGE, transfer to PVDF membrane, and probe with modification-specific antibody (e.g., anti-phospho-(substrate motif)).
- Option B (Radioactive/LC-MS): For higher sensitivity, use [γ-32P]ATP or non-radioactive ATP followed by mass spectrometry to detect mass shift.
- Quantification: Measure band intensity (Western) or peak area (MS) to calculate initial velocity and, if performing a substrate concentration series, derive kinetic parameters (Km, Vmax).
Table 1: Comparative Metrics of Orthogonal Validation Methods
| Method | Throughput | Key Measured Parameter | Typical Time Required | Cost per Sample | Information Gained |
|---|---|---|---|---|---|
| Cellular Assay | Medium | Phenotypic score, Pathway activity (fold-change) | 3-7 days | Medium-High | Functional relevance, cellular context, pathway placement |
| SPR | Low | Binding Affinity (KD in nM), Kinetics (ka, kd) | 1-2 days per ligand | High | Direct interaction, biophysical characterization, binding stoichiometry |
| Enzymatic Activity | Medium-High | Reaction Velocity (nmol/min), Catalytic efficiency (kcat/Km) | 1 day | Low-Medium | Direct biochemical function, catalytic parameters, substrate specificity |
Table 2: Example Data from a SIESTA-Identified Kinase-Substrate Pair Validation
| Validation Method | Experimental Condition | Result | Positive Validation Criteria Met? |
|---|---|---|---|
| SIESTA (Primary Screen) | Lysate + Candidate Peptide | ΔTm = +4.2°C | Initial Hit (ΔTm > 2°C) |
| Cellular Assay | WT Kinase vs. CD Mutant transfection | 5.3-fold increase in substrate phosphorylation | Yes (WT-specific effect) |
| SPR | Kinase injected over immobilized substrate | KD = 18.3 ± 2.1 nM | Yes (High-affinity binding) |
| Enzymatic Activity | In vitro kinase reaction, measured by MS | Vmax = 12.7 pmol/min, Km(substrate) = 2.1 µM | Yes (Catalytically competent) |
Visualizations
Title: Orthogonal Validation Workflow After SIESTA Screen
Title: From Thermal Shift to Functional Validation Pathways
Thesis Context: This document details the application notes and experimental protocols for benchmarking the performance of the System-wide Identification of Enzyme Substrates by Thermal Analysis (SIESTA) platform. Robust characterization of sensitivity, throughput, and technical reproducibility is foundational for its application in large-scale, drug-target interaction mapping and substrate discovery research.
1. Introduction to Performance Metrics in SIESTA The SIESTA method infers target engagement and substrate identification by measuring ligand-induced shifts in protein thermal stability. To validate findings for drug development, three core performance metrics must be quantified: Sensitivity (minimum detectable ligand concentration or melting temperature shift), Throughput (number of samples processed per unit time), and Technical Reproducibility (inter- and intra-assay variance). The following protocols and data provide a framework for this essential benchmarking.
2. Key Research Reagent Solutions Table 1: Essential Materials for SIESTA Benchmarking
| Reagent/Material | Function in Benchmarking |
|---|---|
| Reference Ligands (e.g., Staurosporine, ATP analogs) | Well-characterized binders to target kinases/proteins; provide positive controls for thermal shift magnitude (ΔTm). |
| Thermostable Protein Standard (e.g., ThermoLuc) | Fluorescent reporter unaffected by test conditions; controls for well-to-well detection variance. |
| SYPRO Orange Dye | Environment-sensitive fluorophore; binds hydrophobic patches exposed during protein unfolding. |
| Standardized Protein Lysate | A consistent source of target protein (e.g., recombinant, cell lysate) for reproducibility assays. |
| 384-Well Clear PCR Plates | Standardized format for high-throughput thermal profiling in real-time PCR instruments. |
| qPCR Instrument with Thermal Gradient | Enables precise temperature ramping and simultaneous fluorescence monitoring of multiple samples. |
3. Experimental Protocols
Protocol 3.1: Determining Limit of Detection (Sensitivity) Objective: To establish the minimum ligand concentration that produces a statistically significant ΔTm. Method:
- Prepare a dilution series of the reference ligand (e.g., 10 µM to 100 pM, 1:3 serial dilutions) in assay buffer.
- In a 384-well plate, combine:
- 10 µL of protein solution (target at fixed concentration, e.g., 1 µM).
- 10 µL of ligand dilution or buffer control.
- 5 µL of SYPRO Orange dye (diluted as per manufacturer).
- Centrifuge plate, then run thermal melt protocol on qPCR instrument: 25°C to 95°C with 0.5°C increments and fluorescence read at each step.
- Analyze data by plotting fluorescence derivative (-dF/dT) vs. temperature. Fit curves to determine Tm for each condition.
- Plot ΔTm (Tmligand - Tmcontrol) against log[ligand]. Fit sigmoidal dose-response curve. The Limit of Detection (LoD) is defined as the concentration yielding a ΔTm equal to three times the standard deviation of the vehicle control replicates.
Protocol 3.2: High-Throughput Workflow Validation Objective: To maximize sample processing capacity without compromising data quality. Method:
- Plate Layout Design: Utilize a pre-defined layout with interspersed controls (negative, positive, ThermoLuc standard) across the plate to monitor spatial bias.
- Automated Liquid Handling: Use a liquid handler to dispense protein, dye, and ligand stocks. Validate dispensing accuracy via gravimetric analysis and control well fluorescence uniformity.
- Parallel Thermal Cycling: Run full 384-well plates using the instrument's maximum simultaneous capacity.
- Automated Data Pipeline: Export raw fluorescence data for automated processing via script (e.g., Python/R) that performs baseline correction, Tm calculation, and ΔTm output, minimizing manual intervention.
Protocol 3.3: Assessing Technical Reproducibility Objective: To quantify inter-assay, intra-assay, and inter-operator variance. Method:
- Intra-assay Precision: Repeat Protocol 3.1 with a single mid-range ligand concentration (n=16 replicates) on one plate. Calculate coefficient of variation (CV%) for the resulting ΔTm values.
- Inter-assay Precision: Repeat the same experiment across three independent days, with fresh reagent preparations each day (n=8 replicates per day). Perform one-way ANOVA to assess between-day variance.
- Inter-operator Reproducibility: Have two trained researchers independently prepare and run the same plate layout. Compare ΔTm values for key controls via correlation analysis (Pearson's r > 0.98 target).
4. Benchmarking Data Summary Table 2: Representative Benchmarking Data for SIESTA on Model Kinase ABL1
| Performance Metric | Experimental Condition | Result | Acceptance Criterion |
|---|---|---|---|
| Sensitivity (LoD) | Imatinib titration vs. ABL1 kinase domain | LoD = 0.05 µM (ΔTm = 0.3°C) | LoD ≤ 0.1 µM |
| Throughput | Assay setup + data acquisition | 384 samples in 3 hours | > 100 samples/hour |
| Intra-assay Precision | ΔTm for 1 µM Imatinib (n=16) | CV% = 2.1% | CV% < 5% |
| Inter-assay Precision | ΔTm across 3 days (n=24) | CV% = 3.8% | CV% < 10% |
| Dynamic Range | Max ΔTm for ABL1 with saturating Imatinib | ΔTm_max = +8.5°C | ΔTm_max > 5°C |
5. Visualization of Workflows and Pathways
SIESTA Thermal Melt Assay & Data Analysis Pipeline
Integrated Workflow for SIESTA Performance Benchmarking
Integrating SIESTA with Other Omics Layers for Systems Pharmacology
Application Notes
SIESTA (Systematic Identification of Enzyme Specificity by Thermal Analysis) provides a functional proteomics readout of ligand-induced protein thermal stability shifts across the proteome. When integrated with other omics layers, it enables a systems pharmacology framework for identifying drug targets, off-targets, and mechanisms of action within a functional biological context.
- Target Deconvolution & Mechanism of Action (MoA): Integrating SIESTA data with transcriptomics (RNA-seq) and phosphoproteomics allows for the correlation of direct protein-ligand engagement (SIESTA) with downstream cellular responses. A direct target identified by SIESTA will show thermal stabilization, while its downstream pathway members may show phosphorylation changes, followed by transcriptional alterations.
- Polypharmacology Mapping: SIESTA's unbiased proteome-wide screening is ideal for identifying off-target engagements. These hits can be contextualized with metabolomics data to trace the functional consequences of multi-target engagement on metabolic flux and pathway activity.
- Stratifying Drug Response: Combining SIESTA profiles from different cell lines or patient-derived samples with their corresponding genomic (DNA-seq) and transcriptomic profiles can identify biomarkers predictive of drug response, linking genetic background to direct drug-protein interactions.
Table 1: Multi-Omics Data Correlation in Systems Pharmacology
| Omics Layer | Data Type | SIESTA Integration Purpose | Example Insight |
|---|---|---|---|
| Genomics | Mutations, CNVs | Context for target presence/absence. | Explain lack of SIESTA engagement in a mutated target protein. |
| Transcriptomics | RNA expression levels | Correlate target engagement with gene expression changes. | Distinguish direct stabilization (SIESTA) from indirect transcriptional feedback. |
| Proteomics (Abundance) | Protein expression levels | Normalize thermal shift data; confirm target expression. | Identify if engagement is proportional to protein abundance. |
| Phosphoproteomics | Phosphorylation sites | Link target engagement to immediate signaling rewiring. | Connect kinase stabilization to downstream substrate phosphorylation changes. |
| Metabolomics | Metabolite levels | Functional readout of multi-target engagement on pathways. | Correlate off-target stabilization with metabolite pool alterations. |
Protocols
Protocol 1: Integrated SIESTA-Phosphoproteomics Workflow for MoA Studies
Objective: To identify direct kinase targets and their immediate signaling cascades following drug treatment.
Materials:
- TMTpro 16plex Label Reagent Set (Thermo Fisher)
- Titanium Dioxide (TiO2) Phosphopeptide Enrichment Kit
- Cell line of interest
- Compound of interest vs. DMSO vehicle
- MS-compatible detergent (e.g., RapiGest)
- Trypsin/Lys-C protease mix
- High-pH Reversed-Phase Peptide Fractionation Kit
Procedure:
- Cell Treatment & Lysis: Culture cells in triplicate. Treat with compound or DMSO for a predetermined time (e.g., 1-2 hours). Harvest cells, wash with PBS, and lyse in SIESTA-compatible buffer supplemented with phosphatase and protease inhibitors.
- SIESTA Thermal Profiling: Divide lysate. Subject one aliquot to the standard SIESTA protocol: heat denaturation across a temperature gradient, centrifugation, and digestion of soluble proteins with trypsin.
- Phosphoproteome Sample Preparation: On a separate aliquot of the same lysate, perform protein precipitation and resuspension in denaturing buffer. Reduce, alkylate, and digest proteins with trypsin/Lys-C.
- Phosphopeptide Enrichment: Desalt peptides. Enrich phosphorylated peptides using TiO2 beads according to manufacturer protocol.
- TMT Labeling & Pooling: Label the digested peptides from each SIESTA temperature point and each phosphoproteomics sample with unique TMTpro channels. Pool samples proportionally.
- Fractionation & LC-MS/MS: Fractionate the pooled sample using high-pH reversed-phase chromatography. Analyze fractions by LC-MS/MS on an Orbitrap Eclipse or equivalent.
- Data Analysis:
- SIESTA: Process data with standard software (e.g.,
TPPorMSPurity). Fit melting curves, calculate Tm shifts (ΔTm). Significant stabilizations/destabilizations indicate direct binding. - Phosphoproteomics: Search data against a human database. Quantify TMT reporter ions. Identify significantly changed phosphosites (p < 0.05, fold-change > 1.5).
- Integration: Overlap SIESTA-identified protein targets with upstream kinases or phosphatases of altered phosphosites using pathway analysis tools (Ingenuity, STRING).
- SIESTA: Process data with standard software (e.g.,
SIESTA-Phosphoproteomics Integrated Workflow
Protocol 2: Triangulating SIESTA, Transcriptomics, and Metabolomics for Polypharmacology
Objective: To link multi-target engagement to functional pathway perturbations.
Procedure:
- Parallel Sample Generation: Treat cells (n=6) with compound or vehicle. Harvest at matched time points (e.g., 6h and 24h).
- Aliquot 1 (SIESTA): Lyse in MS-compatible buffer for thermal profiling.
- Aliquot 2 (RNA-seq): Preserve in TRIzol reagent.
- Aliquot 3 (Metabolomics): Quench metabolism with cold methanol/acetonitrile and snap-freeze.
- Multi-Omics Data Acquisition:
- Perform SIESTA as in Protocol 1.
- Extract RNA, prepare libraries, and sequence.
- Extract metabolites and analyze via LC-MS (untargeted metabolomics).
- Integrated Bioinformatics Analysis:
- Identify significant ΔTm proteins (SIESTA), differentially expressed genes (DEGs), and altered metabolites.
- Use joint pathway analysis (e.g., MetaboAnalyst 6.0, OmicsNet) to map all three data types onto KEGG or Reactome pathways.
- Construct a network where SIESTA targets are seeds, connected to DEGs and metabolites via known enzymatic relationships.
Multi-Omics Triangulation for Systems Pharmacology
The Scientist's Toolkit
Table 2: Essential Research Reagents & Materials
| Item | Function in Integrated Workflow |
|---|---|
| TMTpro 16plex Isobaric Label Set | Enables multiplexed quantification of up to 16 samples (multiple temps, replicates, conditions) in a single MS run, crucial for combined SIESTA & proteomics. |
| TiO2 or IMAC Magnetic Beads | For selective enrichment of phosphorylated peptides prior to LC-MS/MS, enabling phosphoproteome analysis. |
| High-pH Reversed-Phase Fractionation Kit | Reduces sample complexity by fractionating peptides before LC-MS/MS, increasing proteome coverage. |
| Phosphatase/Protease Inhibitor Cocktails | Preserves the native proteome and phosphoproteome state during cell lysis for both SIESTA and phospho-enrichment. |
| RapiGest or Similar MS-Compatible Detergent | Aids cell lysis and protein solubilization for SIESTA while being easily cleaved prior to MS analysis. |
| Methanol/Acetonitrile (80%) in Water | Standard solvent for quenching metabolism and extracting polar metabolites for subsequent LC-MS metabolomics. |
| Stable Isotope-labeled Internal Standards (e.g., C13, N15) | Used in metabolomics sample preparation for normalization and quality control of LC-MS runs. |
Conclusion
SIESTA thermal analysis represents a powerful and versatile chemoproteomic platform that moves beyond simple target engagement to enable the system-wide, unbiased identification of protein-substrate interactions and metabolic functions. By mastering its foundational principles (Intent 1), rigorous methodology (Intent 2), and optimization strategies (Intent 3), researchers can reliably deconvolute the complex mechanisms of drug action and cellular metabolism. When validated against complementary techniques (Intent 4), SIESTA data provides high-confidence targets for downstream functional studies. The future of SIESTA lies in its integration with phenotypic screening and multi-omics approaches, promising to accelerate drug discovery by revealing novel therapeutic targets, elucidating polypharmacology, and identifying functional biomarkers for personalized medicine and clinical translation.