Research Articles
Strategies and Solutions for Plasmid Instability in Recombinant Protein Production
This article provides a comprehensive guide for researchers and industry professionals on addressing plasmid instability in recombinant microbial strains.
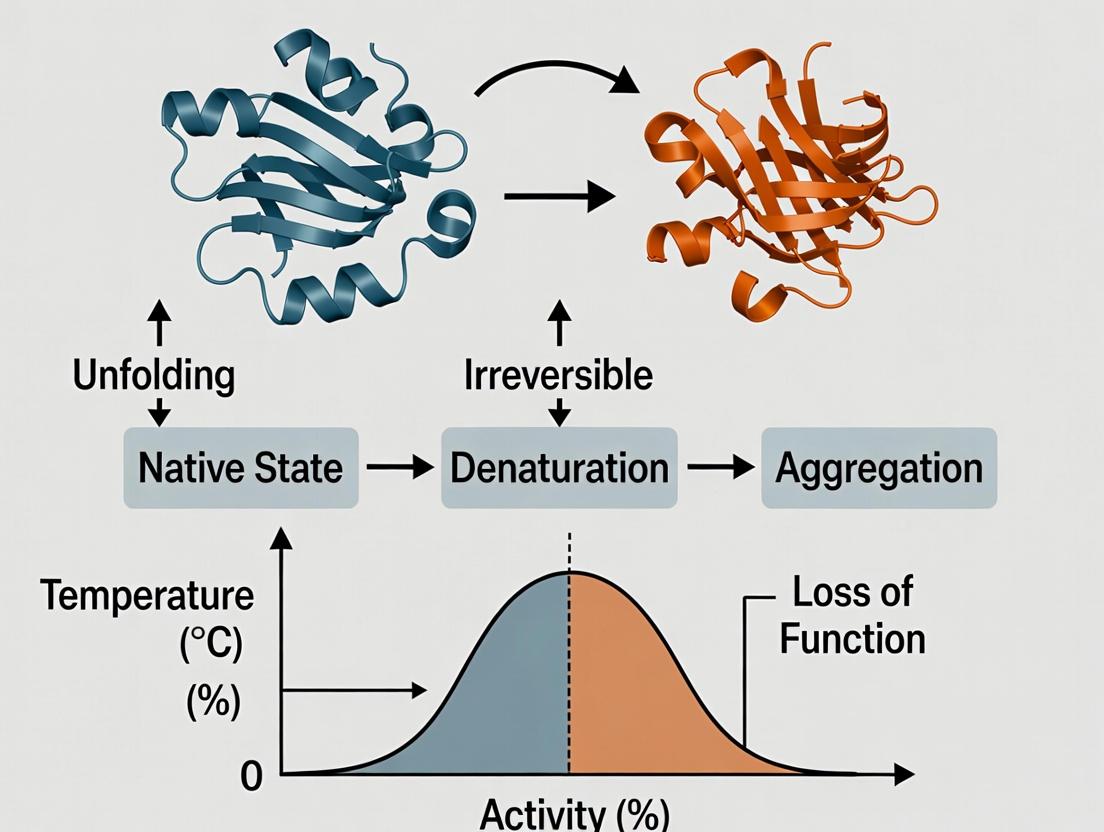
Enzyme Stability Solutions: Advanced Strategies to Prevent Denaturation and Boost Activity in Biotech & Pharma
This article provides a comprehensive guide for researchers and drug development professionals on addressing enzyme instability and denaturation.
Beyond the Cut: Solving Over-Truncation in Enzyme Design for Better Therapeutics
This article addresses the critical challenge of over-truncation in enzyme sequence design for researchers, scientists, and drug development professionals.
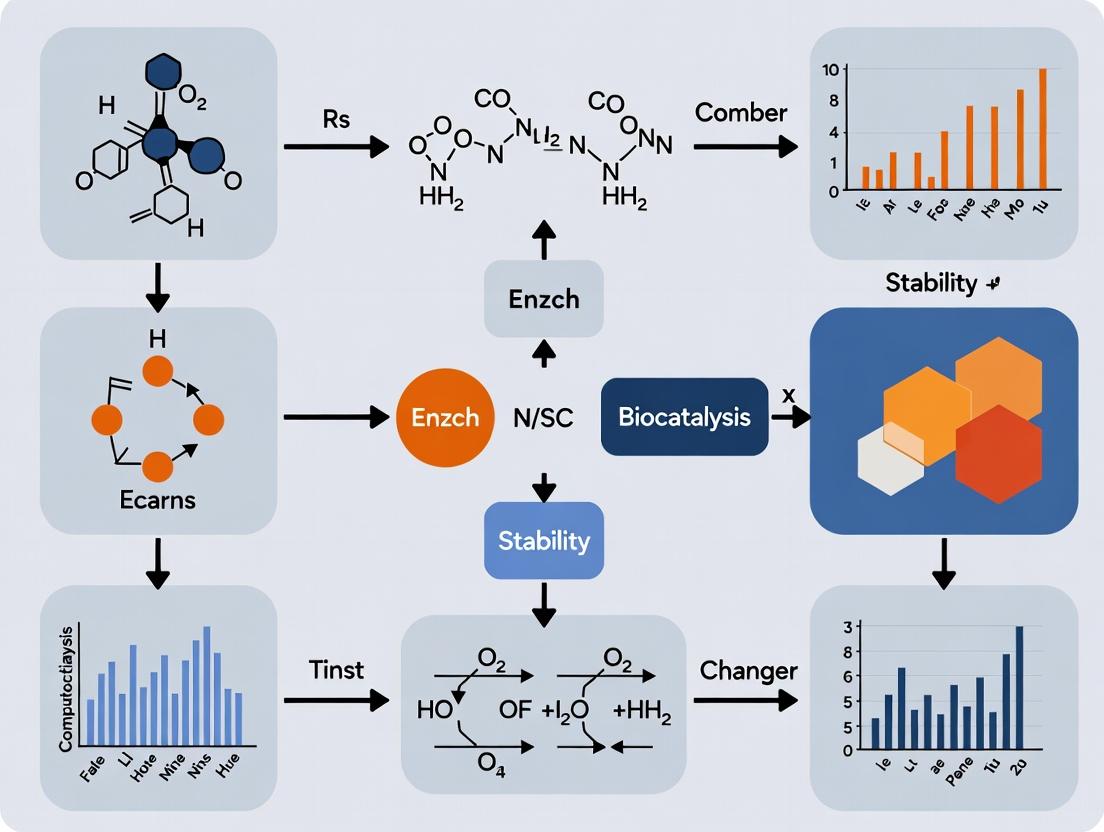
Stabilizing Enzymes in Organic Solvents: Strategies to Overcome Denaturation for Enhanced Biocatalysis and Drug Development
This article provides a comprehensive overview of modern strategies to mitigate organic solvent-induced denaturation in biocatalysis, targeting researchers and pharmaceutical development professionals.
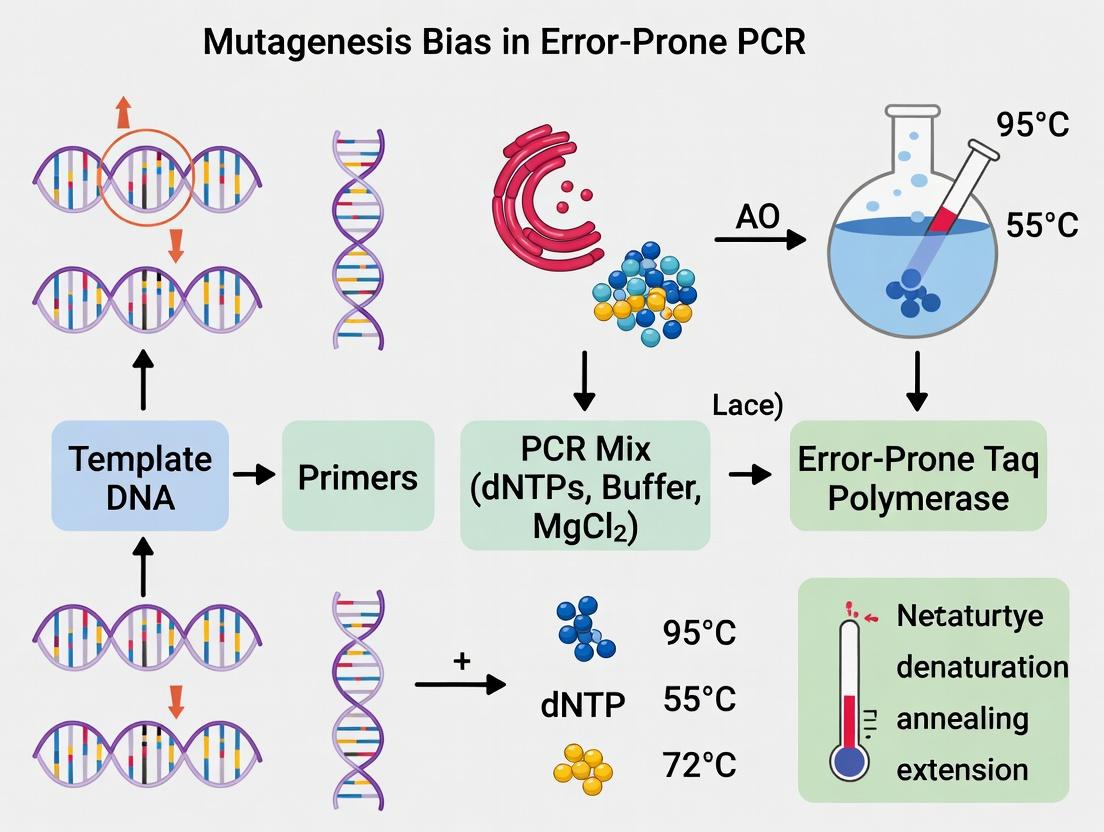
Mastering Error-Prone PCR: Strategies to Overcome Mutagenesis Bias for Directed Evolution Success
This article provides a comprehensive guide for researchers on understanding and mitigating the intrinsic biases of error-prone PCR (epPCR) in directed evolution and protein engineering.
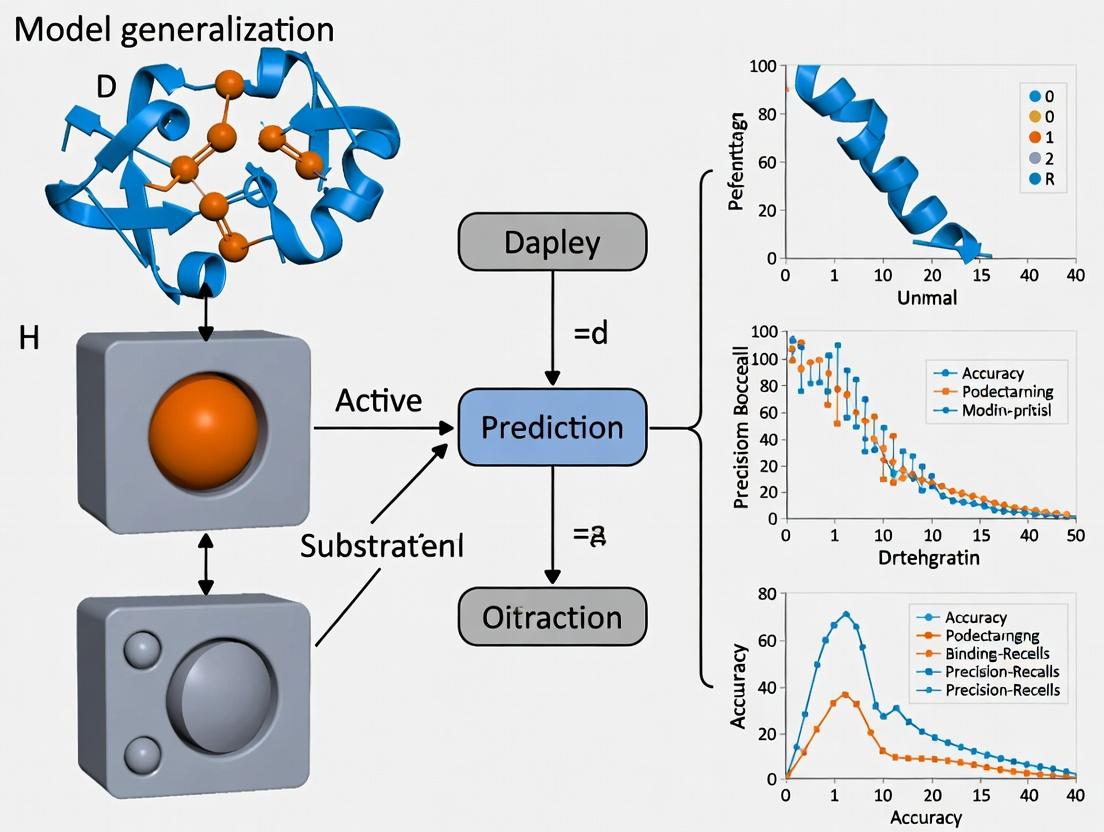
Beyond the Known: Advancing Enzyme Function Prediction Through Enhanced Model Generalization
This article addresses the critical challenge of model generalization in enzyme function prediction, a key bottleneck in translating computational biology to real-world applications.
Taming the Fold: Strategies to Overcome Misfolding in Computationally Designed Enzymes for Biomedical Applications
The promise of computationally designed enzymes for drug development and synthetic biology is frequently hampered by protein misfolding, which leads to aggregation, instability, and loss of function.
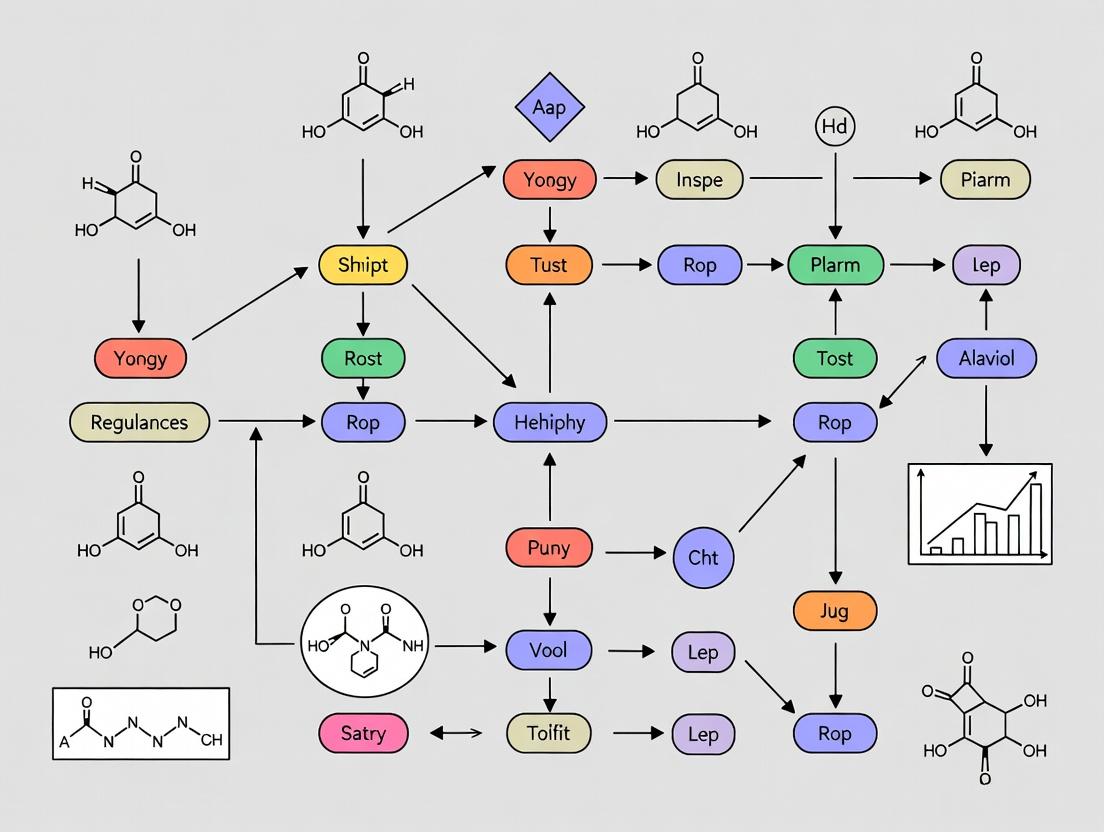
Dynamic Metabolic Networks: Uncovering and Correcting System-Level Imbalances in Disease and Therapeutics
This article provides a comprehensive overview of contemporary strategies for addressing metabolic network imbalances, focusing on their dynamic regulation in biomedical research and drug discovery.
Breaking Through the Bottleneck: Advanced Strategies to Overcome Mass Transfer Limitations in Flow Biocatalysis
This article provides a comprehensive guide for researchers and process engineers on addressing the critical challenge of mass transfer in continuous flow biocatalysis.
Overcoming Mass Transfer Hurdles in Biocatalytic Cascades: Strategies for Enhanced Enzyme Efficiency and Pharmaceutical Synthesis
This article provides a comprehensive guide for researchers and drug development professionals on addressing critical mass transfer limitations in multi-enzyme cascade reactors.